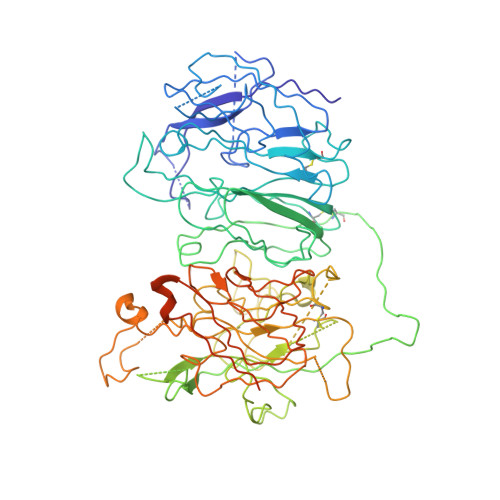
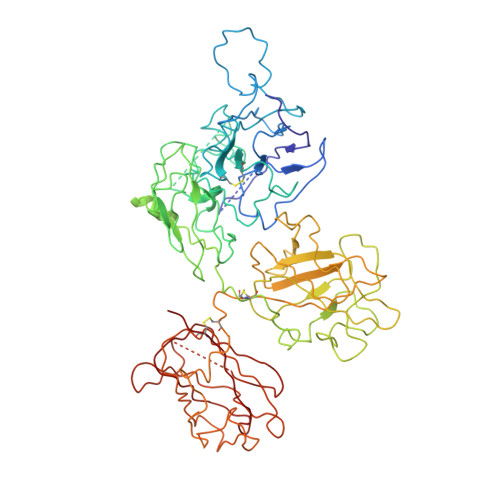
Cryo-EM structures of human coagulation factors V and Va.
Ruben, E.A., Rau, M.J., Fitzpatrick, J.A.J., Di Cera, E.(2021) Blood 137: 3137-3144
- PubMed: 33684942 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1182/blood.2021010684
- Primary Citation Related Structures:
7KVE, 7KVF, 7KXY - PubMed Abstract:
Coagulation factor V (fV) is the precursor of fVa, which, together with fXa, Ca2+, and phospholipids, defines the prothrombinase complex and activates prothrombin in the penultimate step of the coagulation cascade. We solved the cryogenic electron microscopy (cryo-EM) structures of human fV and fVa at atomic (3.3 Å) and near-atomic (4.4 Å) resolution, respectively. The structure of fV reveals the entire A1-A2-B-A3-C1-C2 assembly, but with a surprisingly disordered B domain. The C1 and C2 domains provide a platform for interaction with phospholipid membranes and support the A1 and A3 domains, with the A2 domain sitting on top of them. The B domain is highly dynamic and visible only for short segments connecting to the A2 and A3 domains. The A2 domain reveals all sites of proteolytic processing by thrombin and activated protein C, a partially buried epitope for binding fXa, and fully exposed epitopes for binding activated protein C and prothrombin. Removal of the B domain and activation to fVa exposes the sites of cleavage by activated protein C at R306 and R506 and produces increased disorder in the A1-A2-A3-C1-C2 assembly, especially in the C-terminal acidic portion of the A2 domain that is responsible for prothrombin binding. Ordering of this region and full exposure of the fXa epitope emerge as necessary steps in the assembly of the prothrombin-prothrombinase complex. These structures offer molecular context for the function of fV and fVa and pioneer the analysis of coagulation factors by cryo-EM.
- Edward A. Doisy Department of Biochemistry and Molecular Biology, Saint Louis University School of Medicine, St Louis, MO.
Organizational Affiliation: