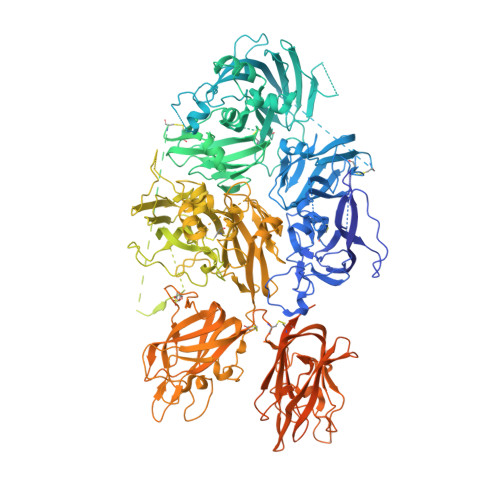
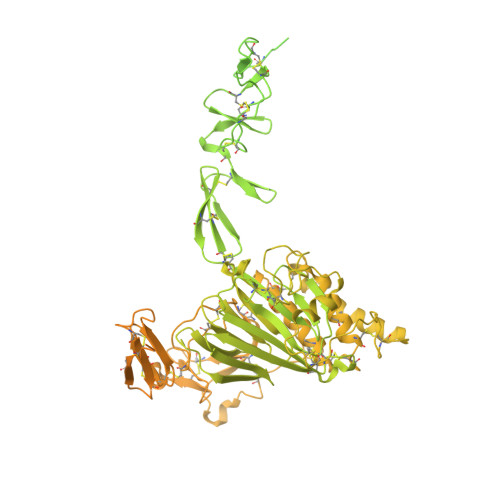
Molecular determinants of the factor VIII/von Willebrand factor complex revealed by BIVV001 cryo-electron microscopy.
Fuller, J.R., Knockenhauer, K.E., Leksa, N.C., Peters, R.T., Batchelor, J.D.(2021) Blood 137: 2970-2980
- PubMed: 33569592 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1182/blood.2020009197
- Primary Citation Related Structures:
7KWO - PubMed Abstract:
Interaction of factor VIII (FVIII) with von Willebrand factor (VWF) is mediated by the VWF D'D3 domains and thrombin-mediated release is essential for hemostasis after vascular injury. VWF-D'D3 mutations resulting in loss of FVIII binding are the underlying cause of von Willebrand disease (VWD) type 2N. Furthermore, the FVIII-VWF interaction has significant implications for the development of therapeutics for bleeding disorders, particularly hemophilia A, in which endogenous VWF clearance imposes a half-life ceiling on replacement FVIII therapy. To understand the structural basis of FVIII engagement by VWF, we solved the structure of BIVV001 by cryo-electron microscopy to 2.9 Å resolution. BIVV001 is a bioengineered clinical-stage FVIII molecule for the treatment of hemophilia A. In BIVV001, VWF-D'D3 is covalently linked to an Fc domain of a B domain-deleted recombinant FVIII (rFVIII) Fc fusion protein, resulting in a stabilized rFVIII/VWF-D'D3 complex. Our rFVIII/VWF structure resolves BIVV001 architecture and provides a detailed spatial understanding of previous biochemical and clinical observations related to FVIII-VWF engagement. Notably, the FVIII acidic a3 peptide region (FVIII-a3), established as a critical determinant of FVIII/VWF complex formation, inserts into a basic groove formed at the VWF-D'/rFVIII interface. Our structure shows direct interaction of sulfated Y1680 in FVIII-a3 and VWF-R816 that, when mutated, leads to severe hemophilia A or VWD type 2N, respectively. These results provide insight on this key coagulation complex, explain the structural basis of many hemophilia A and VWD type 2N mutations, and inform studies to further elucidate how VWF dissociates rapidly from FVIII upon activation.
- Integrated Drug Discovery, Sanofi, Waltham, MA.
Organizational Affiliation: