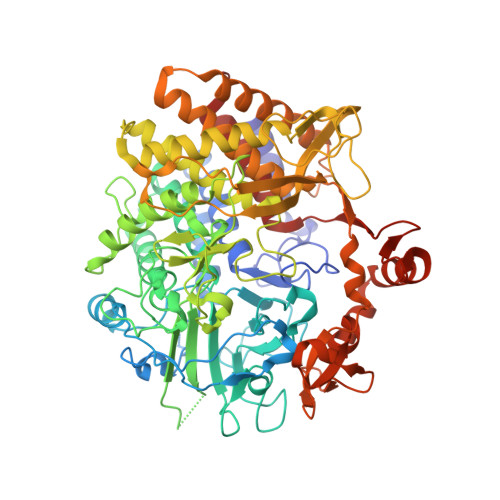
Impact of cellulose properties on enzymatic degradation by bacterial GH48 enzymes: Structural and mechanistic insights from processive Bacillus licheniformis Cel48B cellulase.
Araujo, E.A., Dias, A.H.S., Kadowaki, M.A.S., Piyadov, V., Pellegrini, V.O.A., Urio, M.B., Ramos, L.P., Skaf, M.S., Polikarpov, I.(2021) Carbohydr Polym 264: 118059-118059
- PubMed: 33910709 Search on PubMed
- DOI: https://doi.org/10.1016/j.carbpol.2021.118059
- Primary Citation Related Structures:
7KW6 - PubMed Abstract:
Processive cellulases are highly efficient molecular engines involved in the cellulose breakdown process. However, the mechanism that processive bacterial enzymes utilize to recruit and retain cellulose strands in the catalytic site remains poorly understood. Here, integrated enzymatic assays, protein crystallography and computational approaches were combined to study the enzymatic properties of the processive BlCel48B cellulase from Bacillus licheniformis. Hydrolytic efficiency, substrate binding affinity, cleavage patterns, and the apparent processivity of bacterial BlCel48B are significantly impacted by the cellulose size and its surface morphology. BlCel48B crystallographic structure was solved with ligands spanning -5 to -2 and +1 to +2 subsites. Statistical coupling analysis and molecular dynamics show that co-evolved residues on active site are critical for stabilizing ligands in the catalytic tunnel. Our results provide mechanistic insights into BlCel48B molecular-level determinants of activity, substrate binding, and processivity on insoluble cellulose, thus shedding light on structure-activity correlations of GH48 family members in general.
- São Carlos Institute of Physics, University of São Paulo (USP), São Carlos 13560-970, São Paulo, Brazil; Brazilian Synchrotron Light Laboratory (LNLS), Brazilian Center for Research in Energy and Materials, Campinas 13083-970, São Paulo, Brazil.
Organizational Affiliation: