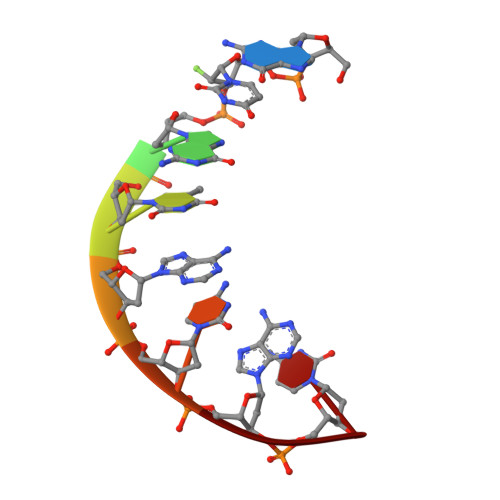
Synthesis and Structural Characterization of 2'-Deoxy-2'-fluoro-l-uridine Nucleic Acids.
Dantsu, Y., Zhang, Y., Zhang, W.(2021) Org Lett 23: 5007-5011
- PubMed: 34142829 Search on PubMed
- DOI: https://doi.org/10.1021/acs.orglett.1c01498
- Primary Citation Related Structures:
7KW4 - PubMed Abstract:
Despite the development of artificial l-RNA/DNA as therapeutic molecules, the in-depth investigation on their chemical modifications is still limited. Here, we synthesize a chemically derivatized 2'-deoxy-2'-fluoro-l-uridine building block and incorporate it into oligonucleotides. Our thermo-denaturization and enzymatic digestion experiments reveal their superior stability. Furthermore, one crystal structure of l-type fluoro-DNA is determined to characterize its handedness. Our results reveal the increase of l-helix stability by fluoro-modification and provide the foundation for its future functional application.
- Department of Biochemistry and Molecular Biology, Indiana University School of Medicine, Indianapolis, Indiana 46202, United States.
Organizational Affiliation: