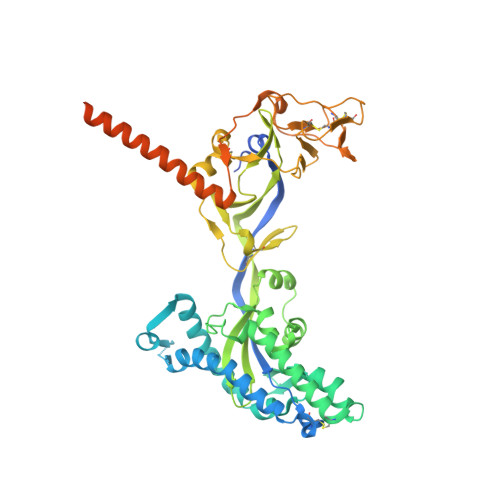
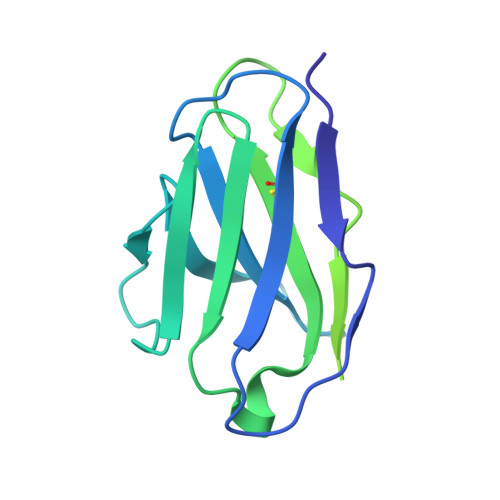
Broadly neutralizing antibody cocktails targeting Nipah virus and Hendra virus fusion glycoproteins.
Dang, H.V., Cross, R.W., Borisevich, V., Bornholdt, Z.A., West, B.R., Chan, Y.P., Mire, C.E., Da Silva, S.C., Dimitrov, A.S., Yan, L., Amaya, M., Navaratnarajah, C.K., Zeitlin, L., Geisbert, T.W., Broder, C.C., Veesler, D.(2021) Nat Struct Mol Biol 28: 426-434
- PubMed: 33927387 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-021-00584-8
- Primary Citation Related Structures:
7KI4, 7KI6 - PubMed Abstract:
Hendra virus (HeV) and Nipah virus (NiV) are henipaviruses (HNVs) causing respiratory illness and severe encephalitis in humans, with fatality rates of 50-100%. There are no licensed therapeutics or vaccines to protect humans. HeV and NiV use a receptor-binding glycoprotein (G) and a fusion glycoprotein (F) to enter host cells. HNV F and G are the main targets of the humoral immune response, and the presence of neutralizing antibodies is a correlate of protection against NiV and HeV in experimentally infected animals. We describe here two cross-reactive F-specific antibodies, 1F5 and 12B2, that neutralize NiV and HeV through inhibition of membrane fusion. Cryo-electron microscopy structures reveal that 1F5 and 12B2 recognize distinct prefusion-specific, conserved quaternary epitopes and lock F in its prefusion conformation. We provide proof-of-concept for using antibody cocktails for neutralizing NiV and HeV and define a roadmap for developing effective countermeasures against these highly pathogenic viruses.
- Department of Biochemistry, University of Washington, Seattle, WA, USA.
Organizational Affiliation: