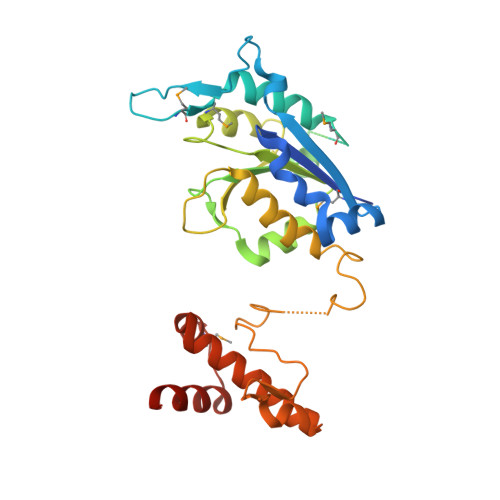
Crystal structure of TrmD tRNA (guanine-N1)-methyltransferase from Corynebacterium diphtheriae in complex with SAH
Michalska, K., Tanase, L., Maltseva, N., Kim, Y., Endres, M., Joachimiak, A., Center for Structural Genomics of Infectious Diseases (CSGID)To be published.