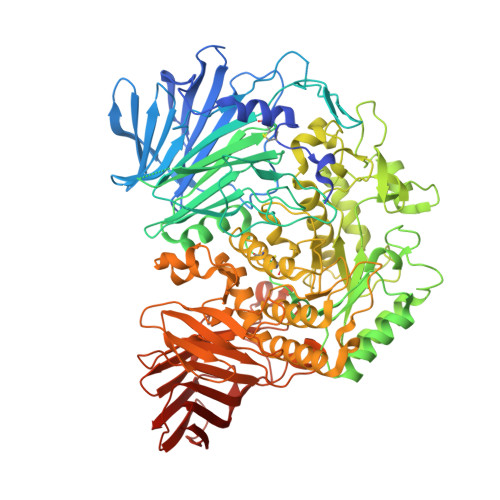
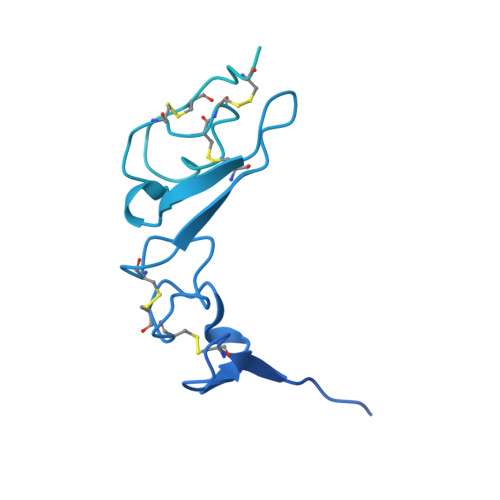
N-Substituted Valiolamine Derivatives as Potent Inhibitors of Endoplasmic Reticulum alpha-Glucosidases I and II with Antiviral Activity.
Karade, S.S., Hill, M.L., Kiappes, J.L., Manne, R., Aakula, B., Zitzmann, N., Warfield, K.L., Treston, A.M., Mariuzza, R.A.(2021) J Med Chem 64: 18010-18024
- PubMed: 34870992 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.1c01377
- Primary Citation Related Structures:
7JTY, 7K9N, 7K9O, 7K9Q, 7K9T, 7KAD, 7KB6, 7KB8, 7KBJ, 7KBR, 7KRY, 7L9E - PubMed Abstract:
Most enveloped viruses rely on the host cell endoplasmic reticulum (ER) quality control (QC) machinery for proper folding of glycoproteins. The key ER α-glucosidases (α-Glu) I and II of the ERQC machinery are attractive targets for developing broad-spectrum antivirals. Iminosugars based on deoxynojirimycin have been extensively studied as ER α-glucosidase inhibitors; however, other glycomimetic compounds are less established. Accordingly, we synthesized a series of N-substituted derivatives of valiolamine, the iminosugar scaffold of type 2 diabetes drug voglibose. To understand the basis for up to 100,000-fold improved inhibitory potency, we determined high-resolution crystal structures of mouse ER α-GluII in complex with valiolamine and 10 derivatives. The structures revealed extensive interactions with all four α-GluII subsites. We further showed that N-substituted valiolamines were active against dengue virus and SARS-CoV-2 in vitro . This study introduces valiolamine-based inhibitors of the ERQC machinery as candidates for developing potential broad-spectrum therapeutics against the existing and emerging viruses.
- University of Maryland Institute for Bioscience and Biotechnology Research, Rockville, Maryland 20850, United States.
Organizational Affiliation: