Discovery of a Novel and Brain-Penetrant O -GlcNAcase Inhibitor via Virtual Screening, Structure-Based Analysis, and Rational Lead Optimization.
Tawada, M., Fushimi, M., Masuda, K., Sun, H., Uchiyama, N., Kosugi, Y., Lane, W., Tjhen, R., Endo, S., Koike, T.(2021) J Med Chem 64: 1103-1115
- PubMed: 33404239 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.0c01712
- Primary Citation Related Structures:
7K41 - PubMed Abstract:
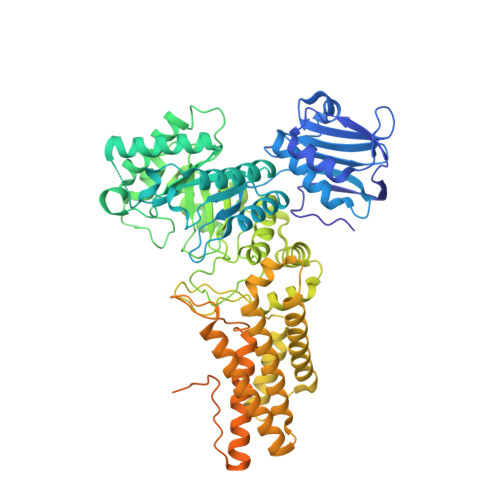
O -GlcNAcase (OGA) has received increasing attention as an attractive therapeutic target for tau-mediated neurodegenerative disorders; however, its role in these pathologies remains unclear. Therefore, potent chemical tools with favorable pharmacokinetic profiles are desirable to characterize this enzyme. Herein, we report the discovery of a potent and novel OGA inhibitor, compound 5i , comprising an aminopyrimidine scaffold, identified by virtual screening based on multiple methodologies combining structure-based and ligand-based approaches, followed by sequential optimization with a focus on ligand lipophilicity efficiency. This compound was observed to increase the level of O -GlcNAcylated protein in cells and display suitable pharmacokinetic properties and brain permeability. Crystallographic analysis revealed that the chemical series bind to OGA via characteristic hydrophobic interactions, which resulted in a high affinity for OGA with moderate lipophilicity. Compound 5i could serve as a useful chemical probe to help establish a proof-of-concept of OGA inhibition as a therapeutic target for the treatment of tauopathies.
- Research, Takeda Pharmaceutical Company Limited, 26-1 Muraoka-Higashi, 2-Chome, Fujisawa, Kanagawa 251-8555, Japan.
Organizational Affiliation: