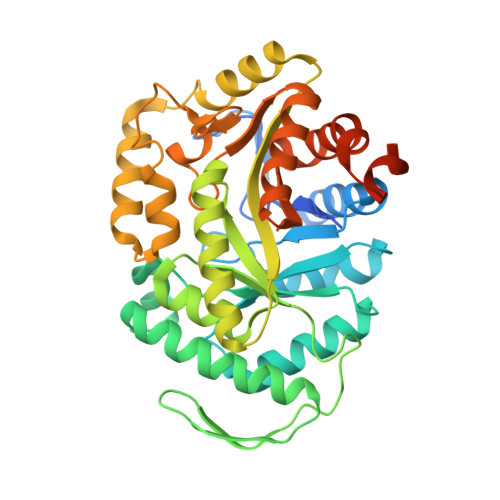
Structures of the alkanesulfonate monooxygenase MsuD provide insight into C-S bond cleavage, substrate scope, and an unexpected role for the tetramer.
Liew, J.J.M., El Saudi, I.M., Nguyen, S.V., Wicht, D.K., Dowling, D.P.(2021) J Biological Chem 297: 100823-100823
- PubMed: 34029591 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2021.100823
- Primary Citation Related Structures:
7JV3, 7JW9, 7JYB, 7K14, 7K64 - PubMed Abstract:
Bacterial two-component flavin-dependent monooxygenases cleave the stable C-S bond of environmental and anthropogenic organosulfur compounds. The monooxygenase MsuD converts methanesulfonate (MS - ) to sulfite, completing the sulfur assimilation process during sulfate starvation, but the mechanism of this conversion remains unclear. To explore the mechanism of C-S bond cleavage, we report a series of crystal structures of MsuD from Pseudomonas fluorescens in different liganded states. This report provides the first crystal structures of an alkanesulfonate monooxygenase with a bound flavin and alkanesulfonate, elucidating the roles of the active site lid, the protein C terminus, and an active site loop in flavin and/or alkanesulfonate binding. These structures position MS - closest to the flavin N5 position, consistent with an N5-(hydro)peroxyflavin mechanism rather than a classical C4a-(hydro)peroxyflavin mechanism. A fully enclosed active site is observed in the ternary complex, mediated by interchain interaction of the C terminus at the tetramer interface. These structures identify an unexpected function of the protein C terminus in this protein family in stabilizing tetramer formation and the alkanesulfonate-binding site. Spurred by interest from the crystal structures, we conducted biochemical assays and molecular docking that redefine MsuD as a small- to medium-chain alkanesulfonate monooxygenase. Functional mutations verify the sulfonate-binding site and reveal the critical importance of the protein C terminus for monooxygenase function. These findings reveal a deeper understanding of MsuD's functionality at the molecular level and consequently how it operates within its role as part of the sulfur assimilation pathway.
- Department of Chemistry, University of Massachusetts Boston, Boston, Massachusetts, USA.
Organizational Affiliation: