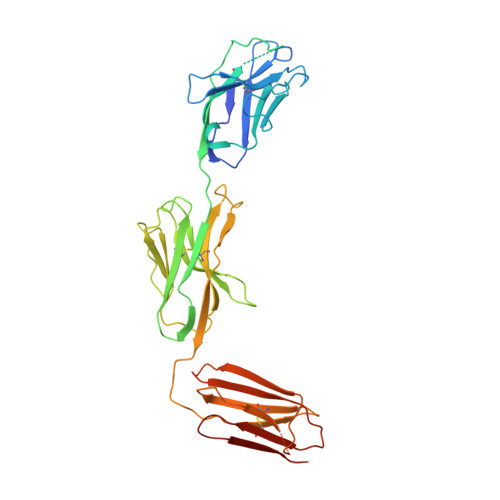
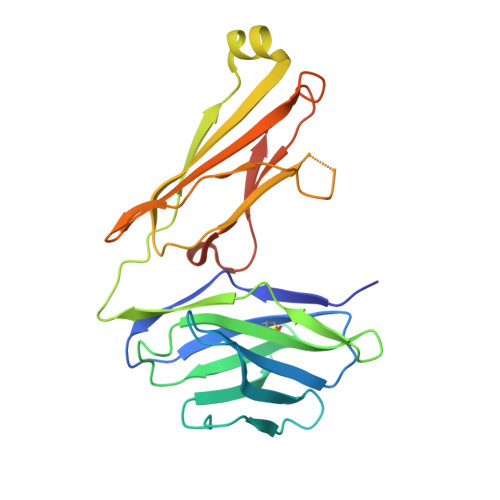
The molecular assembly of the marsupial gamma mu T cell receptor defines a third T cell lineage.
Morrissey, K.A., Wegrecki, M., Praveena, T., Hansen, V.L., Bu, L., Sivaraman, K.K., Darko, S., Douek, D.C., Rossjohn, J., Miller, R.D., Le Nours, J.(2021) Science 371: 1383-1388
- PubMed: 33766885 Search on PubMed
- DOI: https://doi.org/10.1126/science.abe7070
- Primary Citation Related Structures:
7K0X, 7K0Z, 7L15 - PubMed Abstract:
αβ and γδ T cell receptors (TCRs) are highly diverse antigen receptors that define two evolutionarily conserved T cell lineages. We describe a population of γμTCRs found exclusively in non-eutherian mammals that consist of a two-domain (Vγ-Cγ) γ-chain paired to a three-domain (Vμ-Vμj-Cμ) μ-chain. γμTCRs were characterized by restricted diversity in the Vγ and Vμj domains and a highly diverse unpaired Vμ domain. Crystal structures of two distinct γμTCRs revealed the structural basis of the association of the γμTCR heterodimer. The Vμ domain shared the characteristics of a single-domain antibody within which the hypervariable CDR3μ loop suggests a major antigen recognition determinant. We define here the molecular basis underpinning the assembly of a third TCR lineage, the γμTCR.
- Department of Biology, Center for Evolutionary and Theoretical Immunology, University of New Mexico, Albuquerque, NM, USA.
Organizational Affiliation: