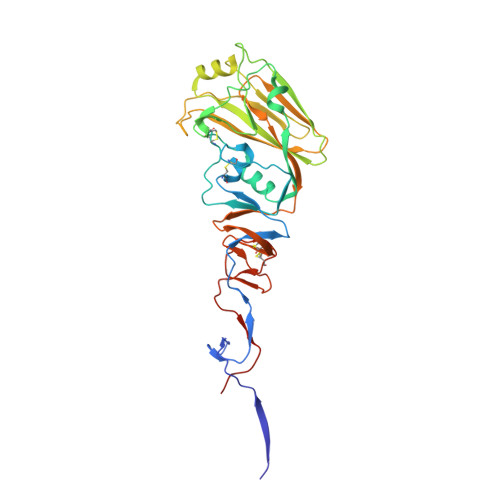Structural characterisation of hemagglutinin from seven Influenza A H1N1 strains reveal diversity in the C05 antibody recognition site.
Ghafoori, S.M., Petersen, G.F., Conrady, D.G., Calhoun, B.M., Stigliano, M.Z.Z., Baydo, R.O., Grice, R., Abendroth, J., Lorimer, D.D., Edwards, T.E., Forwood, J.K.(2023) Sci Rep 13: 6940-6940
- PubMed: 37117205 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-023-33529-w
- Primary Citation Related Structures:
6D8W, 6ML8, 6MYA, 6N41, 6ONA, 6OSR, 7JP4, 7JPD - PubMed Abstract:
Influenza virus (IV) causes several outbreaks of the flu each year resulting in an economic burden to the healthcare system in the billions of dollars. Several influenza pandemics have occurred during the last century and estimated to have caused 100 million deaths. There are four genera of IV, A (IVA), B (IVB), C (IVC), and D (IVD), with IVA being the most virulent to the human population. Hemagglutinin (HA) is an IVA surface protein that allows the virus to attach to host cell receptors and enter the cell. Here we have characterised the high-resolution structures of seven IVA HAs, with one in complex with the anti-influenza head-binding antibody C05. Our analysis revealed conserved receptor binding residues in all structures, as seen in previously characterised IV HAs. Amino acid conservation is more prevalent on the stalk than the receptor binding domain (RBD; also called the head domain), allowing the virus to escape from antibodies targeting the RBD. The equivalent site of C05 antibody binding to A/Denver/57 HA appears hypervariable in the other H1N1 IV HAs. Modifications within this region appear to disrupt binding of the C05 antibody, as these HAs no longer bind the C05 antibody by analytical SEC. Our study brings new insights into the structural and functional recognition of IV HA proteins and can contribute to further development of anti-influenza vaccines.
- School of Dentistry and Medical Sciences, Charles Sturt University, Wagga Wagga, NSW, 2650, Australia.
Organizational Affiliation: