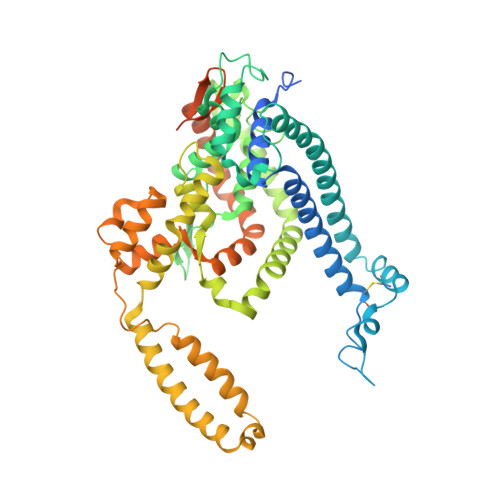
Structure, lipid scrambling activity and role in autophagosome formation of ATG9A.
Maeda, S., Yamamoto, H., Kinch, L.N., Garza, C.M., Takahashi, S., Otomo, C., Grishin, N.V., Forli, S., Mizushima, N., Otomo, T.(2020) Nat Struct Mol Biol 27: 1194-1201
- PubMed: 33106659 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-020-00520-2
- Primary Citation Related Structures:
7JLO, 7JLP, 7JLQ - PubMed Abstract:
De novo formation of the double-membrane compartment autophagosome is seeded by small vesicles carrying membrane protein autophagy-related 9 (ATG9), the function of which remains unknown. Here we find that ATG9A scrambles phospholipids of membranes in vitro. Cryo-EM structures of human ATG9A reveal a trimer with a solvated central pore, which is connected laterally to the cytosol through the cavity within each protomer. Similarities to ABC exporters suggest that ATG9A could be a transporter that uses the central pore to function. Moreover, molecular dynamics simulation suggests that the central pore opens laterally to accommodate lipid headgroups, thereby enabling lipids to flip. Mutations in the pore reduce scrambling activity and yield markedly smaller autophagosomes, indicating that lipid scrambling by ATG9A is essential for membrane expansion. We propose ATG9A acts as a membrane-embedded funnel to facilitate lipid flipping and to redistribute lipids added to the outer leaflet of ATG9 vesicles, thereby enabling growth into autophagosomes.
- Department of Integrative Structural and Computational Biology, The Scripps Research Institute, La Jolla, CA, USA.
Organizational Affiliation: