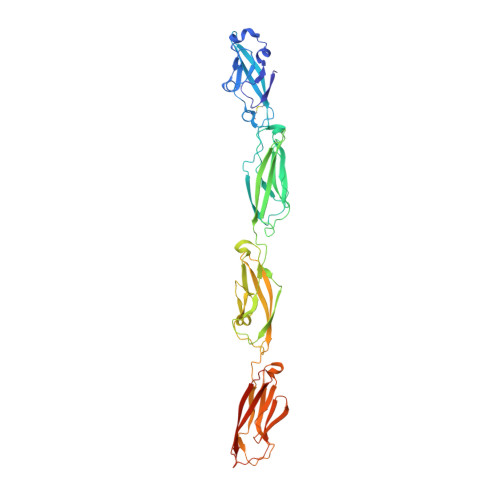How clustered protocadherin binding specificity is tuned for neuronal self-/nonself-recognition.
Goodman, K.M., Katsamba, P.S., Rubinstein, R., Ahlsen, G., Bahna, F., Mannepalli, S., Dan, H., Sampogna, R.V., Shapiro, L., Honig, B.(2022) Elife 11
- PubMed: 35253643 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.72416
- Primary Citation Related Structures:
7JGZ, 7RGF - PubMed Abstract:
The stochastic expression of fewer than 60 clustered protocadherin (cPcdh) isoforms provides diverse identities to individual vertebrate neurons and a molecular basis for self-/nonself-discrimination. cPcdhs form chains mediated by alternating cis and trans interactions between apposed membranes, which has been suggested to signal self-recognition. Such a mechanism requires that cPcdh cis dimers form promiscuously to generate diverse recognition units, and that trans interactions have precise specificity so that isoform mismatches terminate chain growth. However, the extent to which cPcdh interactions fulfill these requirements has not been definitively demonstrated. Here, we report biophysical experiments showing that cPcdh cis interactions are promiscuous, but with preferences favoring formation of heterologous cis dimers. Trans homophilic interactions are remarkably precise, with no evidence for heterophilic interactions between different isoforms. A new C-type cPcdh crystal structure and mutagenesis data help to explain these observations. Overall, the interaction characteristics we report for cPcdhs help explain their function in neuronal self-/nonself-discrimination.
- Zuckerman Mind, Brain and Behavior Institute, Columbia University, New York, United States.
Organizational Affiliation: