The cross-reaction complex structure with VQIFNK peptide and the tau antibody's Fab domain.
Tsuchida, T., Tsuchiya, T., Miyamoto, K., In, Y., Minoura, K., Taniguchi, t., Ishida, T., Tomoo, K.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Fab Heavy Chain | A [auth H] | 219 | Mus musculus | Mutation(s): 0 | 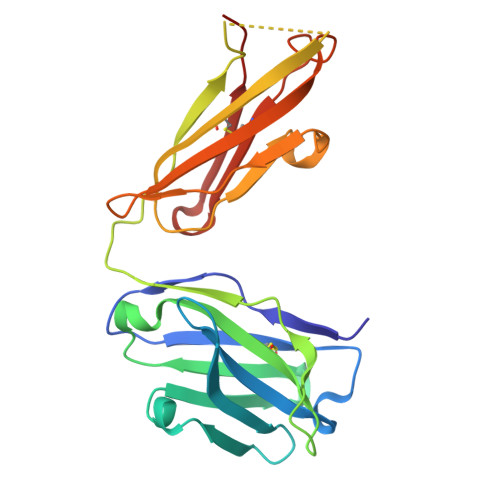 |
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Fab Light Chain | B [auth L] | 217 | Mus musculus | Mutation(s): 0 | 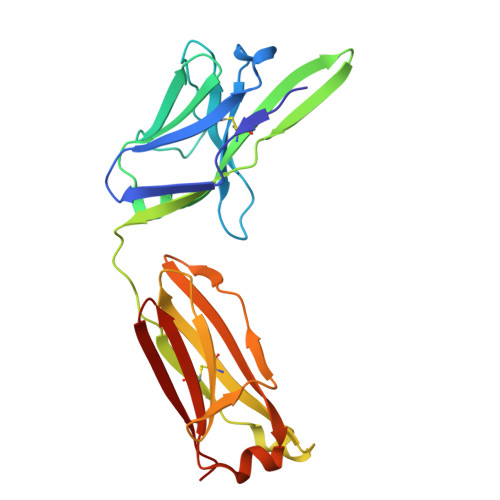 |
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| VQIFNK | 6 | Homo sapiens | Mutation(s): 0 | 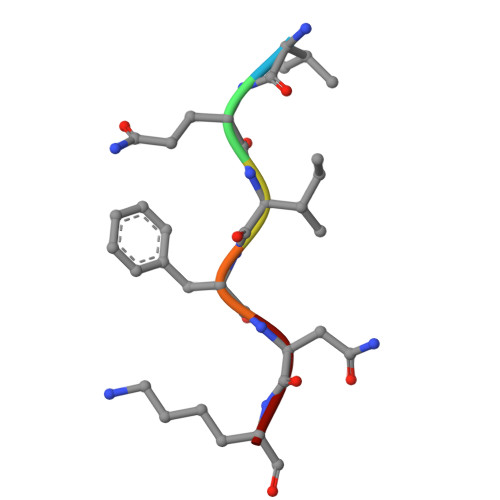 | |
| Ligands 4 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| IOD Download:Ideal Coordinates CCD File | K [auth H] | IODIDE ION I XMBWDFGMSWQBCA-UHFFFAOYSA-M |  | ||
| GOL Download:Ideal Coordinates CCD File | D [auth H], E [auth H], F [auth H], L, M [auth L] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| CL Download:Ideal Coordinates CCD File | I [auth H], J [auth H], N [auth L] | CHLORIDE ION Cl VEXZGXHMUGYJMC-UHFFFAOYSA-M |  | ||
| NH4 Download:Ideal Coordinates CCD File | G [auth H], H | AMMONIUM ION H4 N QGZKDVFQNNGYKY-UHFFFAOYSA-O |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 179.05 | α = 90 |
| b = 70.14 | β = 93.15 |
| c = 41.26 | γ = 90 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| CrystalClear | data reduction |
| CrystalClear | data scaling |
| MOLREP | phasing |
| Coot | model building |
| PDB_EXTRACT | data extraction |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Not funded | -- |