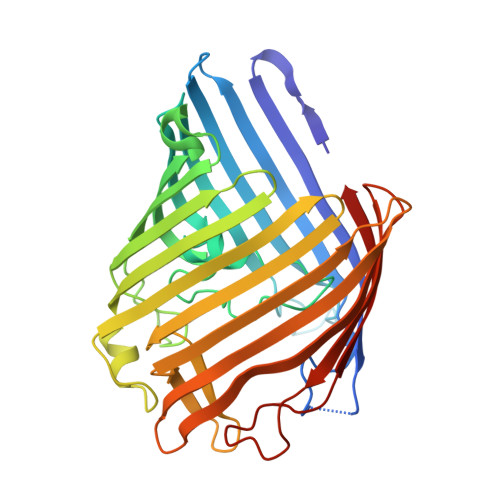
Design and directed evolution of noncanonical beta-stereoselective metalloglycosidases.
Jeong, W.J., Song, W.J.(2022) Nat Commun 13: 6844-6844
- PubMed: 36369431 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-34713-8
- Primary Citation Related Structures:
7FDY, 7FF7 - PubMed Abstract:
Metallohydrolases are ubiquitous in nearly all subclasses of hydrolases, utilizing metal elements to activate a water molecule and facilitate its subsequent dissociation of diverse chemical bonds. However, such a catalytic role of metal ions is rarely found with glycosidases that hydrolyze the glycosidic bonds in sugars. Herein, we design metalloglycosidases by constructing a hydrolytically active Zn-binding site within a barrel-shaped outer membrane protein OmpF. Structure- and mechanism-based redesign and directed evolution have led to the emergence of Zn-dependent glycosidases with catalytic proficiency of 2.8 × 10 9 and high β-stereoselectivity. Biochemical characterizations suggest that the Zn-binding site constitutes a key catalytic motif along with at least one adjacent acidic residue. This work demonstrates that unprecedented metalloenzymes can be tailor-made, expanding the scope of inorganic reactivities in proteinaceous environments, resetting the structural and functional diversity of metalloenzymes, and providing the potential molecular basis of unidentified metallohydrolases and novel whole-cell biocatalysts.
- Department of Chemistry, Seoul National University, Seoul, 08826, Republic of Korea.
Organizational Affiliation: