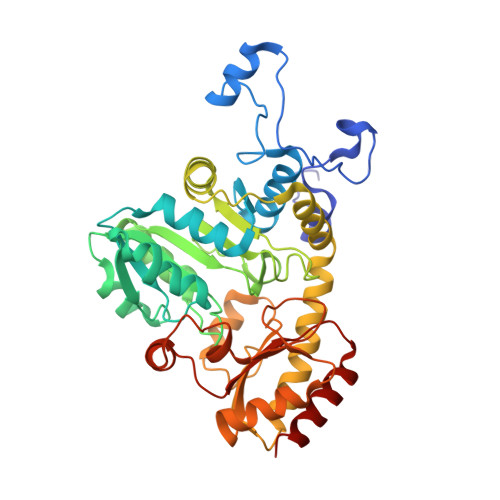
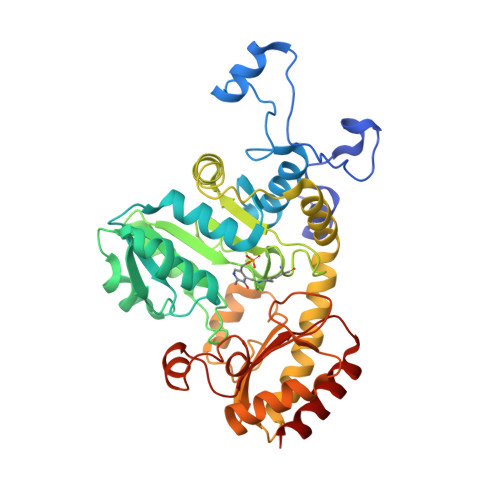
Characterization and application of l-methionine gamma-lyase Q349S mutant enzyme with an enhanced activity toward l-homocysteine.
Okawa, A., Handa, H., Yasuda, E., Murota, M., Kudo, D., Tamura, T., Shiba, T., Inagaki, K.(2022) J Biosci Bioeng 133: 213-221
- PubMed: 34953671 Search on PubMed
- DOI: https://doi.org/10.1016/j.jbiosc.2021.11.008
- Primary Citation Related Structures:
7F1P, 7F1U, 7F1V - PubMed Abstract:
l-Methionine γ-lyse (MGL), a pyridoxal 5'-phosphate-dependent enzyme, catalyzes the α,γ-elimination of l-methionine (l-Met) and l-homocysteine (l-Hcy) to produce α-keto acids, thiols, and ammonia. Previously, various mutant enzymes of Pseudomonas putida MGL (PpMGL) were prepared to identify a homocysteine (Hcy)-specific enzyme that would assist the diagnosis of homocystinuria. Among the mutat enzymes the Q349S mutant exhibited high degradation activity toward l-Hcy. In the present study, PpMGL Q349S was characterized; the results suggested that it could be applied to determine the amount of l-Hcy. Compared to the wild-type PpMGL, specific activities of the Q349S mutant with l-Hcy and l-Met were 1.5 and 0.7 times, respectively. Additionally, we confirmed that l-Hcy in plasma samples could be accurately detected using the Q349S mutant by preincubating it with cysteine desulfurase (CsdA). Furthermore, we determined the X-ray crystal structure of PpMGL Q349S l-Met or l-Hcy complexes Michaelis complex, germinal diamine, and external aldimine at 2.25-2.40 Å. These 3D structures showed that the interaction partner of the β-hydroxyl group of Thr355 in the wild-type PpMGL was changed to the carboxyl group of the Hcy-PLP external aldimine in the Q349S mutant. The interaction of Ser349 and Arg375 was different between l-Met and l-Hcy recognition, indicating that it was important for the recognition of the carboxyl group of the substrate.
- Department of Biofunctional Chemistry, Graduate School of Environmental and Life Science, Okayama University, Okayama 700-8530, Japan. Electronic address: pn2b9nr4@s.okayama-u.ac.jp.
Organizational Affiliation: