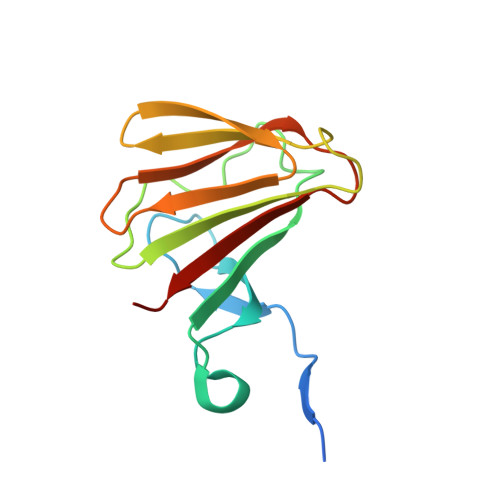
Structural and Mechanistic Bases for StnK3 and Its Mutant-Mediated Lewis-Acid-Dependent Epimerization and Retro-Aldol Reactions.
Chen, M.H., Li, Y.S., Hsu, N.S., Lin, K.H., Wang, Y.L., Wang, Z.C., Chang, C.F., Lin, J.P., Chang, C.Y., Li, T.L.(2022) ACS Catal 12: 1945-1956