Activation and allosteric regulation of the orphan GPR88-Gi1 signaling complex.
Chen, G., Xu, J., Inoue, A., Schmidt, M.F., Bai, C., Lu, Q., Gmeiner, P., Liu, Z., Du, Y.(2022) Nat Commun 13: 2375-2375
- PubMed: 35501348 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-30081-5
- Primary Citation Related Structures:
7EJX, 7WZ4 - PubMed Abstract:
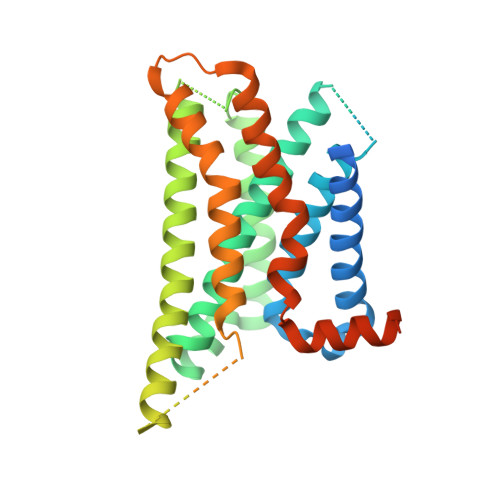
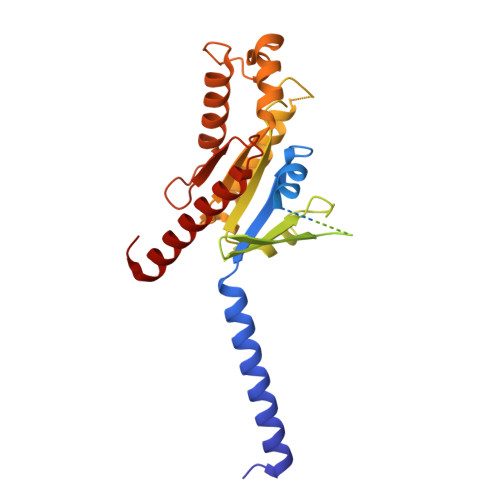
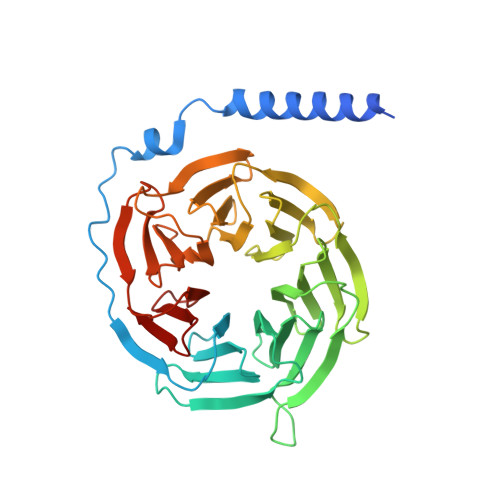
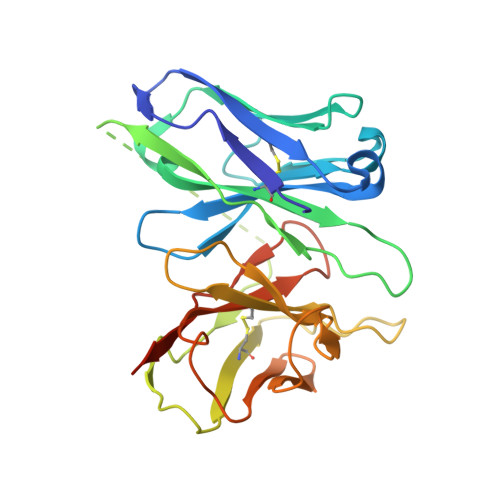
GPR88 is an orphan class A G-protein-coupled receptor that is highly expressed in the striatum and regulates diverse brain and behavioral functions. Here we present cryo-EM structures of the human GPR88-Gi1 signaling complex with or without a synthetic agonist (1R, 2R)-2-PCCA. We show that (1R, 2R)-2-PCCA is an allosteric modulator binding to a herein identified pocket formed by the cytoplasmic ends of transmembrane segments 5, 6, and the extreme C terminus of the α5 helix of Gi1. We also identify an electron density in the extracellular orthosteric site that may represent a putative endogenous ligand of GPR88. These structures, together with mutagenesis studies and an inactive state model obtained from metadynamics simulations, reveal a unique activation mechanism for GPR88 with a set of distinctive structure features and a water-mediated polar network. Overall, our results provide a structural framework for understanding the ligand binding, activation and signaling mechanism of GPR88, and will facilitate the innovative drug discovery for neuropsychiatric disorders and for deorphanization of this receptor.
- Kobilka Institute of Innovative Drug Discovery, School of Medicine, Chinese University of Hong Kong, Shenzhen, Guangdong, 518172, China.
Organizational Affiliation: