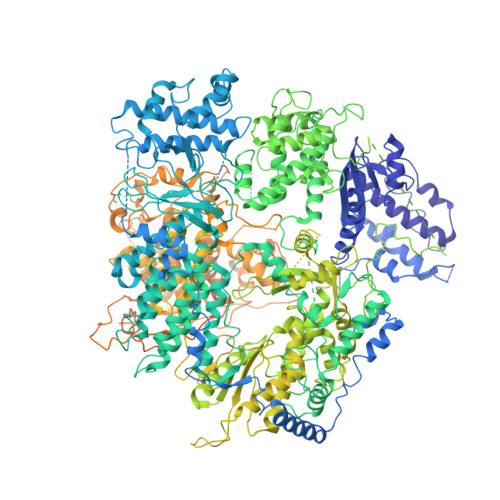
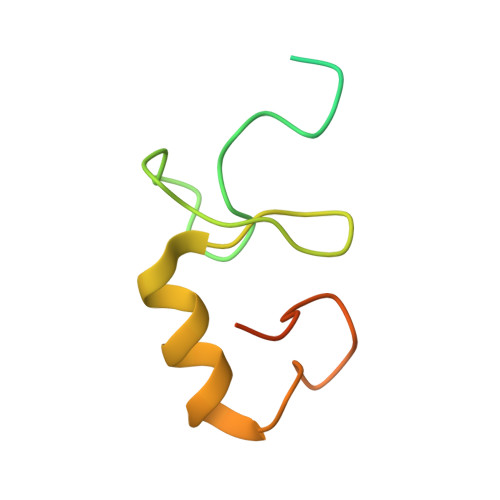
Structural basis for recognition and regulation of arenavirus polymerase L by Z protein.
Kang, H., Cong, J., Wang, C., Ji, W., Xin, Y., Qian, Y., Li, X., Chen, Y., Rao, Z.(2021) Nat Commun 12: 4134-4134
- PubMed: 34226547 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-24458-1
- Primary Citation Related Structures:
7EJU - PubMed Abstract:
Junin virus (JUNV) causes Argentine hemorrhagic fever, a debilitating human disease of high mortality rates and a great risk to public health worldwide. Studying the L protein that replicates and transcribes the genome of JUNV, and its regulator Z protein should provide critical clues to identify therapeutic targets for disrupting the life cycle of JUNV. Here we report the 3.54 Å cryo-EM structure of the JUNV L protein complexed with regulator Z protein. JUNV L structure reveals a conserved architecture containing signature motifs found in other L proteins. Structural analysis shows that L protein is regulated by binding of Z protein at the RNA product exit site. Based on these findings, we propose a model for the role of Z protein as a switch to turn on/off the viral RNA synthesis via its interaction with L protein. Our work unveils the mechanism of JUNV transcription, replication and regulation, which provides a framework for the rational design of antivirals for combating viral infections.
- National Laboratory of Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, Beijing, China.
Organizational Affiliation: