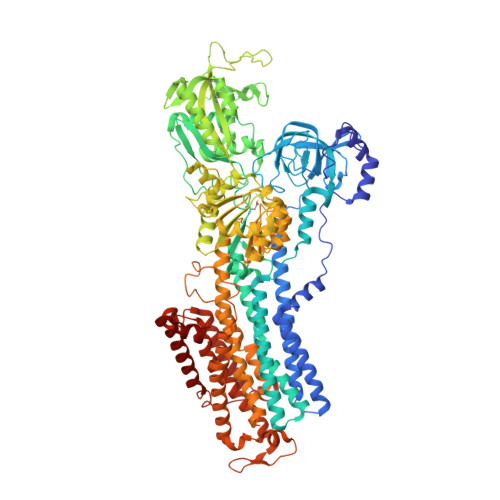
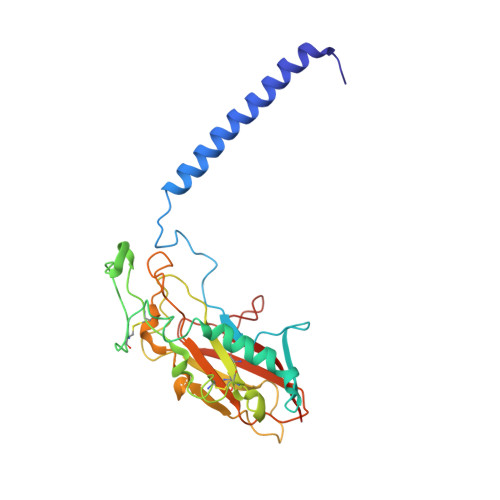
Gastric proton pump with two occluded K + engineered with sodium pump-mimetic mutations.
Abe, K., Yamamoto, K., Irie, K., Nishizawa, T., Oshima, A.(2021) Nat Commun 12: 5709-5709
- PubMed: 34588453 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-26024-1
- Primary Citation Related Structures:
7EFL, 7EFM, 7EFN, 7ET1 - PubMed Abstract:
The gastric H + ,K + -ATPase mediates electroneutral exchange of 1H + /1K + per ATP hydrolysed across the membrane. Previous structural analysis of the K + -occluded E2-P transition state of H + ,K + -ATPase showed a single bound K + at cation-binding site II, in marked contrast to the two K + ions occluded at sites I and II of the closely-related Na + ,K + -ATPase which mediates electrogenic 3Na + /2K + translocation across the membrane. The molecular basis of the different K + stoichiometry between these K + -counter-transporting pumps is elusive. We show a series of crystal structures and a cryo-EM structure of H + ,K + -ATPase mutants with changes in the vicinity of site I, based on the structure of the sodium pump. Our step-wise and tailored construction of the mutants finally gave a two-K + bound H + ,K + -ATPase, achieved by five mutations, including amino acids directly coordinating K + (Lys791Ser, Glu820Asp), indirectly contributing to cation-binding site formation (Tyr340Asn, Glu936Val), and allosterically stabilizing K + -occluded conformation (Tyr799Trp). This quintuple mutant in the K + -occluded E2-P state unambiguously shows two separate densities at the cation-binding site in its 2.6 Å resolution cryo-EM structure. These results offer new insights into how two closely-related cation pumps specify the number of K + accommodated at their cation-binding site.
- Cellular and Structural Physiology Institute, Nagoya University, Nagoya, 464-8601, Japan. kabe@cespi.nagoya-u.ac.jp.
Organizational Affiliation: