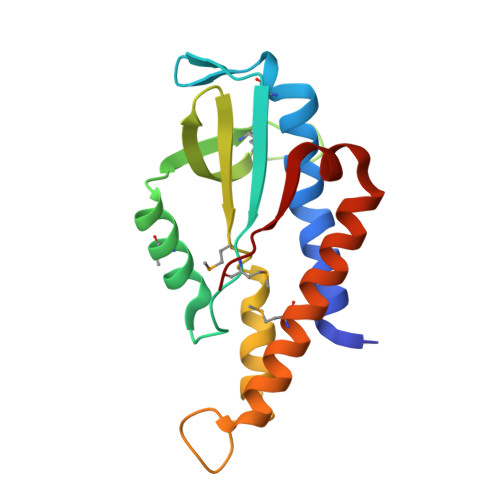
Crystal structure and functional implication of bacterial STING.
Ko, T.-P., Wang, Y.-C., Yang, C.-S., Hou, M.-H., Chen, C.-J., Chiu, Y.-F., Chen, Y.(2022) Nat Commun 13: 26-26
- PubMed: 35013136 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-26583-3
- Primary Citation Related Structures:
7EBD, 7EBL - PubMed Abstract:
Mammalian innate immune sensor STING (STimulator of INterferon Gene) was recently found to originate from bacteria. During phage infection, bacterial STING sense c-di-GMP generated by the CD-NTase (cGAS/DncV-like nucleotidyltransferase) encoded in the same operon and signal suicide commitment as a defense strategy that restricts phage propagation. However, the precise binding mode of c-di-GMP to bacterial STING and the specific recognition mechanism are still elusive. Here, we determine two complex crystal structures of bacterial STING/c-di-GMP, which provide a clear picture of how c-di-GMP is distinguished from other cyclic dinucleotides. The protein-protein interactions further reveal the driving force behind filament formation of bacterial STING. Finally, we group the bacterial STING into two classes based on the conserved motif in β-strand lid, which dictate their ligand specificity and oligomerization mechanism, and propose an evolution-based model that describes the transition from c-di-GMP-dependent signaling in bacteria to 2'3'-cGAMP-dependent signaling in eukaryotes.
- Institute of Biological Chemistry, Academia Sinica, Taipei, 115, Taiwan.
Organizational Affiliation: