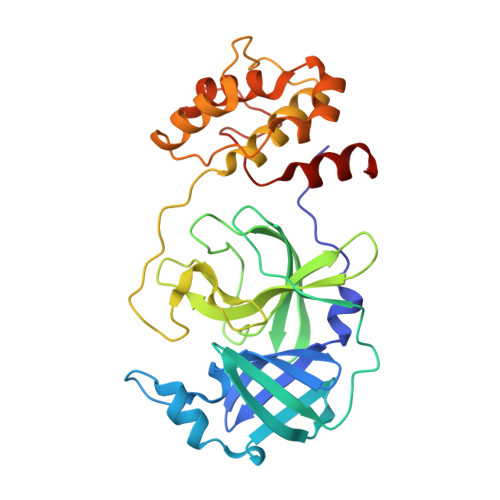
Crystal structures of human coronavirus NL63 main protease at different pH values
Gao, H., Zhang, Y., Jiang, H., Hu, X., Zhang, Y., Zhou, X., Zhong, F., Lin, C., Li, J., Luo, J., Zhang, J.(2021) Acta Crystallogr F Struct Biol Commun 77: 348-355
- PubMed: 34605439 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X21009523
- Primary Citation Related Structures:
7E6L, 7E6M, 7E6N, 7E6R - PubMed Abstract:
Human coronavirus NL63 (HCoV-NL63), which belongs to the genus Alphacoronavirus, mainly infects children and the immunocompromized and is responsible for a series of clinical manifestations, including cough, fever, rhinorrhoea, bronchiolitis and croup. HCoV-NL63, which was first isolated from a seven-month-old child in 2004, has led to infections worldwide and accounts for 10% of all respiratory illnesses caused by etiological agents. However, effective antivirals against HCoV-NL63 infection are currently unavailable. The HCoV-NL63 main protease (M pro ), also called 3C-like protease (3CL pro ), plays a vital role in mediating viral replication and transcription by catalyzing the cleavage of replicase polyproteins (pp1a and pp1ab) into functional subunits. Moreover, M pro is highly conserved among all coronaviruses, thus making it a prominent drug target for antiviral therapy. Here, four crystal structures of HCoV-NL63 M pro in the apo form at different pH values are reported at resolutions of up to 1.78 Å. Comparison with M pro from other human betacoronaviruses such as SARS-CoV-2 and SARS-CoV reveals common and distinct structural features in different genera and extends knowledge of the diversity, function and evolution of coronaviruses.
- Department of Rehabilitation Medicine, The Second Affiliated Hospital of Nanchang University, Nanchang, Jiangxi 330006, People's Republic of China.
Organizational Affiliation: