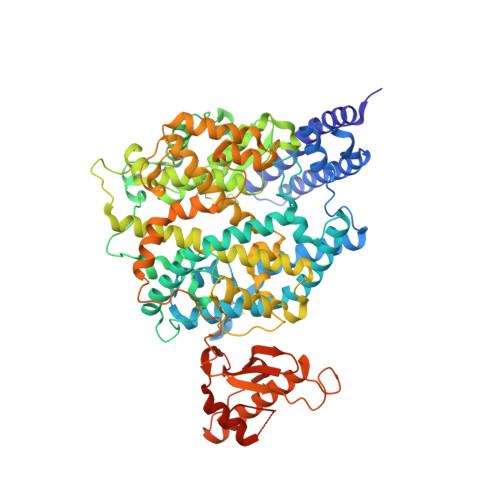
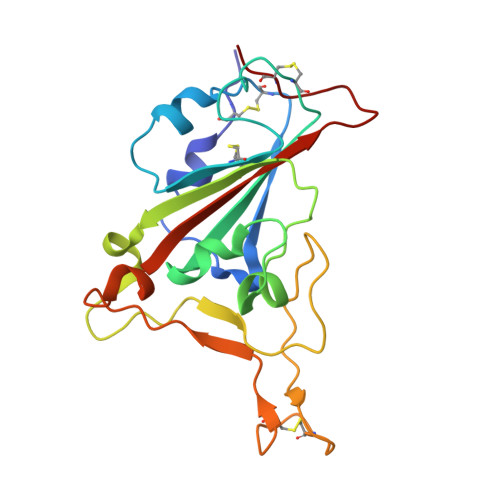
The molecular basis for SARS-CoV-2 binding to dog ACE2.
Zhang, Z., Zhang, Y., Liu, K., Li, Y., Lu, Q., Wang, Q., Zhang, Y., Wang, L., Liao, H., Zheng, A., Ma, S., Fan, Z., Li, H., Huang, W., Bi, Y., Zhao, X., Wang, Q., Gao, G.F., Xiao, H., Tong, Z., Qi, J., Sun, Y.(2021) Nat Commun 12: 4195-4195
- PubMed: 34234119 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-24326-y
- Primary Citation Related Structures:
7E3J - PubMed Abstract:
SARS-CoV-2 can infect many domestic animals, including dogs. Herein, we show that dog angiotensin-converting enzyme 2 (dACE2) can bind to the SARS-CoV-2 spike (S) protein receptor binding domain (RBD), and that both pseudotyped and authentic SARS-CoV-2 can infect dACE2-expressing cells. We solved the crystal structure of RBD in complex with dACE2 and found that the total number of contact residues, contact atoms, hydrogen bonds and salt bridges at the binding interface in this complex are slightly fewer than those in the complex of the RBD and human ACE2 (hACE2). This result is consistent with the fact that the binding affinity of RBD to dACE2 is lower than that of hACE2. We further show that a few important mutations in the RBD binding interface play a pivotal role in the binding affinity of RBD to both dACE2 and hACE2. Our work reveals a molecular basis for cross-species transmission and potential animal spread of SARS-CoV-2, and provides new clues to block the potential transmission chains of this virus.
- CAS Key Laboratory of Pathogenic Microbiology and Immunology, Institute of Microbiology, Chinese Academy of Sciences, Beijing, China.
Organizational Affiliation: