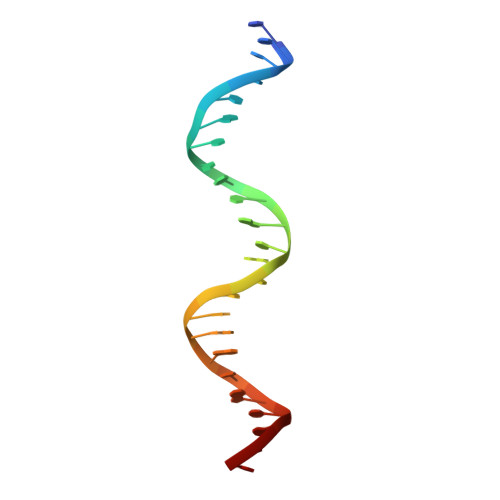
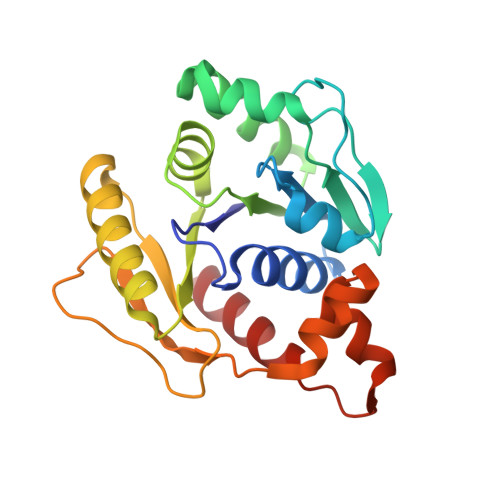
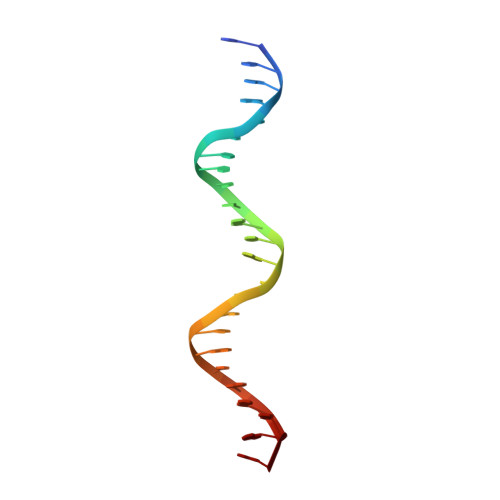
Chromosome segregation in Archaea: SegA- and SegB-DNA complex structures provide insights into segrosome assembly.
Yen, C.Y., Lin, M.G., Chen, B.W., Ng, I.W., Read, N., Kabli, A.F., Wu, C.T., Shen, Y.Y., Chen, C.H., Barilla, D., Sun, Y.J., Hsiao, C.D.(2021) Nucleic Acids Res 49: 13150-13164
- PubMed: 34850144 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkab1155
- Primary Citation Related Structures:
7DUT, 7DUV, 7DV2, 7DV3, 7DWR - PubMed Abstract:
Genome segregation is a vital process in all organisms. Chromosome partitioning remains obscure in Archaea, the third domain of life. Here, we investigated the SegAB system from Sulfolobus solfataricus. SegA is a ParA Walker-type ATPase and SegB is a site-specific DNA-binding protein. We determined the structures of both proteins and those of SegA-DNA and SegB-DNA complexes. The SegA structure revealed an atypical, novel non-sandwich dimer that binds DNA either in the presence or in the absence of ATP. The SegB structure disclosed a ribbon-helix-helix motif through which the protein binds DNA site specifically. The association of multiple interacting SegB dimers with the DNA results in a higher order chromatin-like structure. The unstructured SegB N-terminus plays an essential catalytic role in stimulating SegA ATPase activity and an architectural regulatory role in segrosome (SegA-SegB-DNA) formation. Electron microscopy results also provide a compact ring-like segrosome structure related to chromosome organization. These findings contribute a novel mechanistic perspective on archaeal chromosome segregation.
- Institute of Bioinformatics and Structural Biology, National Tsing Hua University, Hsinchu 300, Taiwan.
Organizational Affiliation: