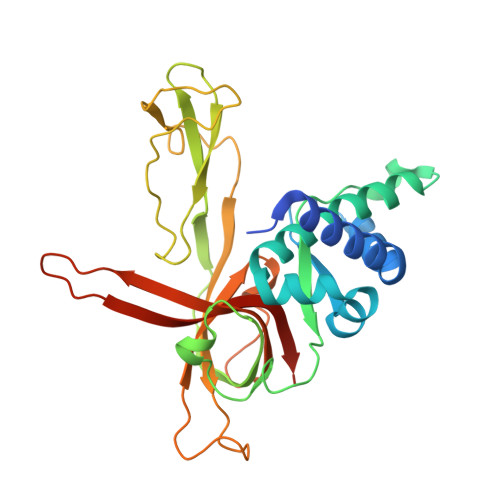
Structural study of the N-terminal domain of human MCM8/9 complex.
Li, J., Yu, D., Liu, L., Liang, H., Ouyang, Q., Liu, Y.(2021) Structure 29: 1171-1181.e4
- PubMed: 34043945 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2021.05.006
- Primary Citation Related Structures:
7DP3, 7DPD - PubMed Abstract:
MCM8/9 is a complex involved in homologous recombination (HR) repair pathway. MCM8/9 dysfunction can cause genome instability and result in primary ovarian insufficiency (POI). However, the mechanism underlying these effects is largely unknown. Here, we report crystal structures of the N-terminal domains (NTDs) of MCM8 and MCM9, and build a ring-shaped NTD structure based on a 6.6 Å resolution cryoelectron microscopy map. This shows that the MCM8/9 complex forms a 3:3 heterohexamer in an alternating pattern. A positively charged DNA binding channel and a putative ssDNA exit pathway for fork DNA unwinding are revealed. Based on the atomic model, the potential effects of the clinical POI mutants are interpreted. Surprisingly, the zinc-finger motifs are found to be capable of binding an iron atom as well. Overall, our results provide a model for the formation of the MCM8/9 complex and provide a path for further studies.
- School of Medicine, Sun Yat-Sen University, Guangdong 510006, China.
Organizational Affiliation: