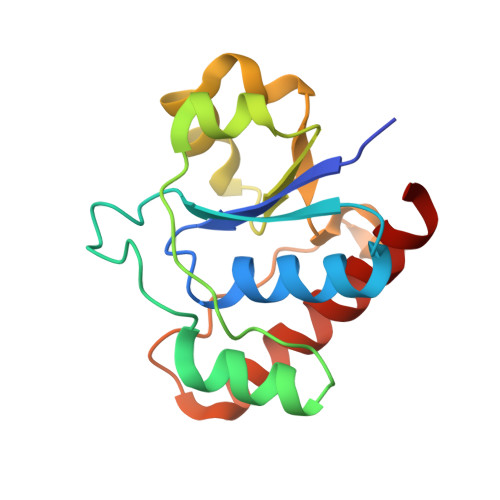
Wzb of Vibrio vulnificus represents a new group of low-molecular-weight protein tyrosine phosphatases with a unique insertion in the W-loop.
Wang, X., Ma, Q.(2021) J Biological Chem 296: 100280-100280
- PubMed: 33450227 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2021.100280
- Primary Citation Related Structures:
7DHD, 7DHE, 7DHF - PubMed Abstract:
Protein tyrosine phosphorylation regulates the production of capsular polysaccharide, an essential virulence factor of the deadly pathogen Vibrio vulnificus. The process requires the protein tyrosine kinase Wzc and its cognate phosphatase Wzb, both of which are largely uncharacterized. Herein, we report the structures of Wzb of V. vulnificus (VvWzb) in free and ligand-bound forms. VvWzb belongs to the low-molecular-weight protein tyrosine phosphatase (LMWPTP) family. Interestingly, it contains an extra four-residue insertion in the W-loop, distinct from all known LMWPTPs. The W-loop of VvWzb protrudes from the protein body in the free structure, but undergoes significant conformational changes to fold toward the active site upon ligand binding. Deleting the four-residue insertion from the W-loop severely impaired the enzymatic activity of VvWzb, indicating its importance for optimal catalysis. However, mutating individual residues or even substituting the whole insertion with four alanine residues only modestly decreased the enzymatic activity, suggesting that the contribution of the insertion to catalysis is not determined by the sequence specificity. Furthermore, inserting the four residues into Escherichia coli Wzb at the corresponding position enhanced its activity as well, indicating that the four-residue insertion in the W-loop can act as a general activity enhancing element for other LMWPTPs. The novel W-loop type and phylogenetic analysis suggested that VvWzb and its homologs should be classified into a new group of LMWPTPs. Our study sheds new insight into the catalytic mechanism and structural diversity of the LMWPTP family and promotes the understanding of the protein tyrosine phosphorylation system in prokaryotes.
- Key Laboratory of Experimental Marine Biology, Institute of Oceanology, Chinese Academy of Sciences, Qingdao, China; Laboratory for Marine Biology and Biotechnology, Pilot National Laboratory for Marine Science and Technology, Qingdao, China; University of Chinese Academy of Sciences, Beijing, China.
Organizational Affiliation: