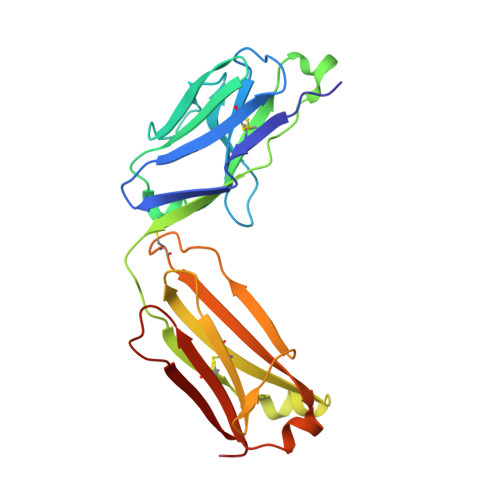
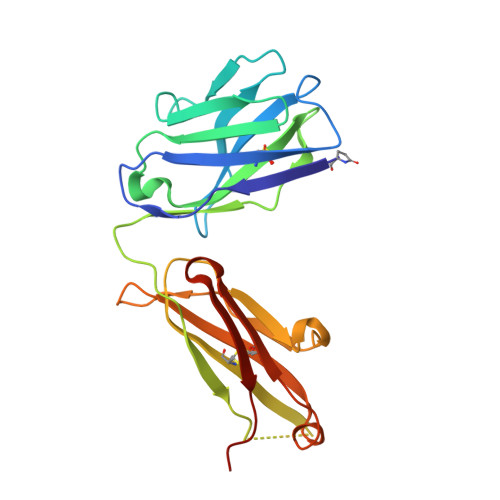
The tertiary structure of the human Xkr8-Basigin complex that scrambles phospholipids at plasma membranes.
Sakuragi, T., Kanai, R., Tsutsumi, A., Narita, H., Onishi, E., Nishino, K., Miyazaki, T., Baba, T., Kosako, H., Nakagawa, A., Kikkawa, M., Toyoshima, C., Nagata, S.(2021) Nat Struct Mol Biol 28: 825-834
- PubMed: 34625749 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-021-00665-8
- Primary Citation Related Structures:
7D9Z, 7DAA, 7DCE - PubMed Abstract:
Xkr8-Basigin is a plasma membrane phospholipid scramblase activated by kinases or caspases. We combined cryo-EM and X-ray crystallography to investigate its structure at an overall resolution of 3.8 Å. Its membrane-spanning region carrying 22 charged amino acids adopts a cuboid-like structure stabilized by salt bridges between hydrophilic residues in transmembrane helices. Phosphatidylcholine binding was observed in a hydrophobic cleft on the surface exposed to the outer leaflet of the plasma membrane. Six charged residues placed from top to bottom inside the molecule were essential for scrambling phospholipids in inward and outward directions, apparently providing a pathway for their translocation. A tryptophan residue was present between the head group of phosphatidylcholine and the extracellular end of the path. Its mutation to alanine made the Xkr8-Basigin complex constitutively active, indicating that it plays a vital role in regulating its scramblase activity. The structure of Xkr8-Basigin provides insights into the molecular mechanisms underlying phospholipid scrambling.
- Laboratory of Biochemistry and Immunology, World Premier International Immunology Frontier Research Center, Osaka University, Suita, Japan.
Organizational Affiliation: