ISRIB Blunts the Integrated Stress Response by Allosterically Antagonising the Inhibitory Effect of Phosphorylated eIF2 on eIF2B.
Zyryanova, A.F., Kashiwagi, K., Rato, C., Harding, H.P., Crespillo-Casado, A., Perera, L.A., Sakamoto, A., Nishimoto, M., Yonemochi, M., Shirouzu, M., Ito, T., Ron, D.(2021) Mol Cell 81: 88
- PubMed: 33220178 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2020.10.031
- Primary Citation Related Structures:
7D43, 7D44, 7D45, 7D46 - PubMed Abstract:
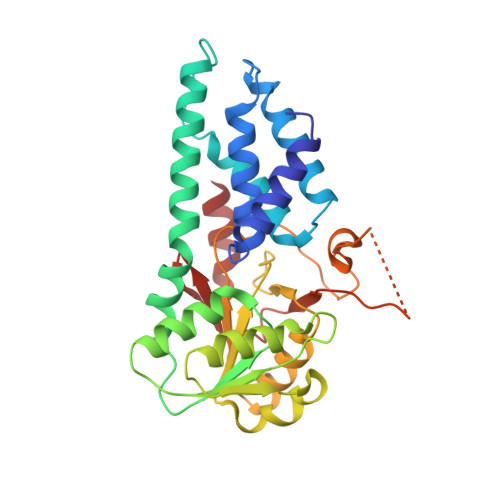
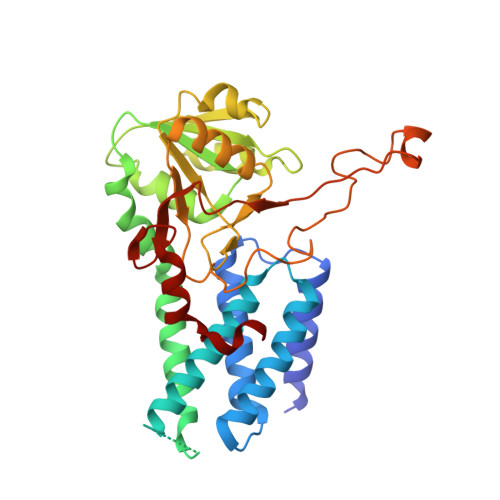
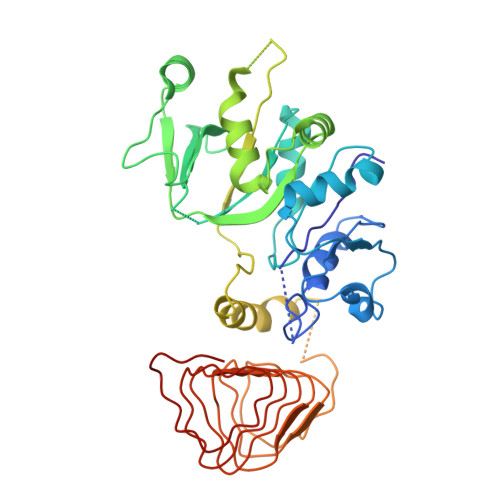
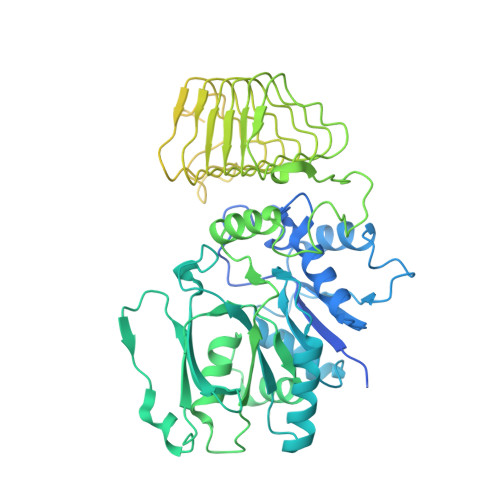
The small molecule ISRIB antagonizes the activation of the integrated stress response (ISR) by phosphorylated translation initiation factor 2, eIF2(αP). ISRIB and eIF2(αP) bind distinct sites in their common target, eIF2B, a guanine nucleotide exchange factor for eIF2. We have found that ISRIB-mediated acceleration of eIF2B's nucleotide exchange activity in vitro is observed preferentially in the presence of eIF2(αP) and is attenuated by mutations that desensitize eIF2B to the inhibitory effect of eIF2(αP). ISRIB's efficacy as an ISR inhibitor in cells also depends on presence of eIF2(αP). Cryoelectron microscopy (cryo-EM) showed that engagement of both eIF2B regulatory sites by two eIF2(αP) molecules remodels both the ISRIB-binding pocket and the pockets that would engage eIF2α during active nucleotide exchange, thereby discouraging both binding events. In vitro, eIF2(αP) and ISRIB reciprocally opposed each other's binding to eIF2B. These findings point to antagonistic allostery in ISRIB action on eIF2B, culminating in inhibition of the ISR.
- Cambridge Institute for Medical Research (CIMR), University of Cambridge, Cambridge CB2 0XY, UK.
Organizational Affiliation: