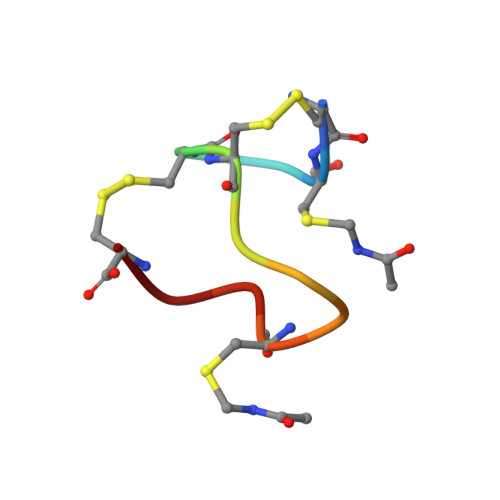
Topological Regulation of the Bioactive Conformation of a Disulfide-Rich Peptide, Heat-Stable Enterotoxin.
Shimamoto, S., Fukutsuji, M., Osumi, T., Goto, M., Toyoda, H., Hidaka, Y.(2020) Molecules 25
- PubMed: 33096591 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/molecules25204798
- Primary Citation Related Structures:
7CSS, 7D37 - PubMed Abstract:
Heat-stable enterotoxin (ST a ) produced by enterotoxigenic E. coli causes acute diarrhea and also can be used as a specific probe for colorectal cancer cells. ST a contains three intra-molecular disulfide bonds (C1-C4, C2-C5, and C3-C6 connectivity). The chemical synthesis of ST a provided not only the native type of ST a but also a topological isomer that had the native disulfide pairings. Interestingly, the activity of the topological isomer was approximately 1/10-1/2 that of the native ST a . To further investigate the bioactive conformation of this molecule and the regulation of disulfide-coupled folding during its chemical syntheses, we examined the folding mechanism of ST a that occurs during its chemical synthesis. The folding intermediate of ST a with two disulfide bonds (C1-C4 and C3-C6) and two Cys(Acm) residues, the precursor peptide, was treated with iodine to produce a third disulfide bond under several conditions. The topological isomer was predominantly produced under all conditions tested, along with trace amounts of the native type of ST a . In addition, NMR measurements indicated that the topological isomer has a left-handed spiral structure similar to that of the precursor peptide, while the native type of ST a had a right-handed spiral structure. These results indicate that the order of the regioselective formation of disulfide bonds is important for the regulation of the final conformation of disulfide-rich peptides in chemical synthesis.
- Faculty of Science and Engineering, Kindai University, 3-4-1 Kowakae, Higashi-Osaka, Osaka 577-8502, Japan.
Organizational Affiliation: