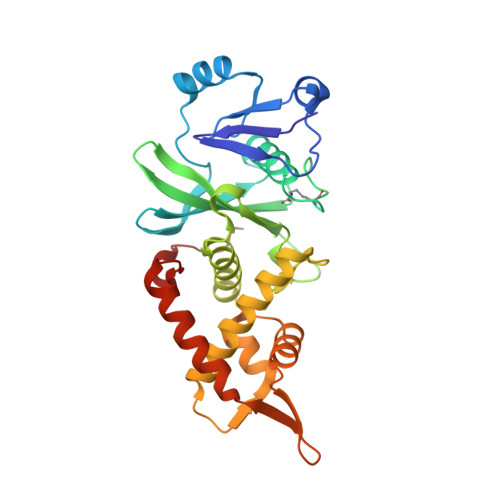
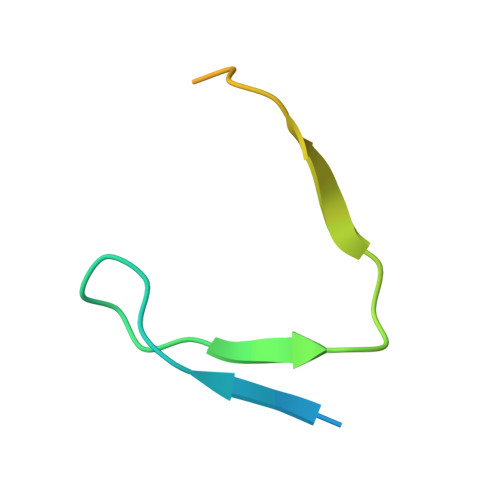
HBO1 is a versatile histone acyltransferase critical for promoter histone acylations.
Xiao, Y., Li, W., Yang, H., Pan, L., Zhang, L., Lu, L., Chen, J., Wei, W., Ye, J., Li, J., Li, G., Zhang, Y., Tan, M., Ding, J., Wong, J.(2021) Nucleic Acids Res 49: 8037-8059
- PubMed: 34259319 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkab607
- Primary Citation Related Structures:
7D0O, 7D0P, 7D0Q, 7D0R, 7D0S - PubMed Abstract:
Recent studies demonstrate that histones are subjected to a series of short-chain fatty acid modifications that is known as histone acylations. However, the enzymes responsible for histone acylations in vivo are not well characterized. Here, we report that HBO1 is a versatile histone acyltransferase that catalyzes not only histone acetylation but also propionylation, butyrylation and crotonylation both in vivo and in vitro and does so in a JADE or BRPF family scaffold protein-dependent manner. We show that the minimal HBO1/BRPF2 complex can accommodate acetyl-CoA, propionyl-CoA, butyryl-CoA and crotonyl-CoA. Comparison of CBP and HBO1 reveals that they catalyze histone acylations at overlapping as well as distinct sites, with HBO1 being the key enzyme for H3K14 acylations. Genome-wide chromatin immunoprecipitation assay demonstrates that HBO1 is highly enriched at and contributes to bulk histone acylations on the transcriptional start sites of active transcribed genes. HBO1 promoter intensity highly correlates with the level of promoter histone acylation, but has no significant correlation with level of transcription. We also show that HBO1 is associated with a subset of DNA replication origins. Collectively our study establishes HBO1 as a versatile histone acyltransferase that links histone acylations to promoter acylations and selection of DNA replication origins.
- Shanghai Key Laboratory of Regulatory Biology, Institute of Biomedical Sciences and School of Life Sciences, East China Normal University, Shanghai 200241, China.
Organizational Affiliation: