Structure of the human CLCN7-OSTM1 complex with ATP
Yang, G.H., Lu, Y.M., Zhang, Y.M., Jia, Y.T., Li, X.M., Lei, J.L.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Osteopetrosis-associated transmembrane protein 1 | 344 | Homo sapiens | Mutation(s): 0 Gene Names: OSTM1, GL, HSPC019, UNQ6098/PRO21201 Membrane Entity: Yes | 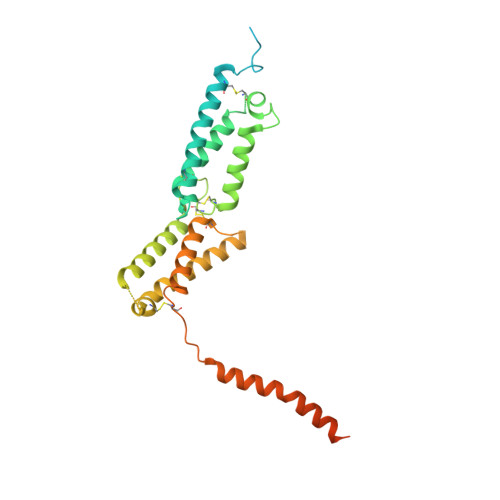 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q86WC4 GTEx: ENSG00000081087 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q86WC4 | ||||
Glycosylation | |||||
| Glycosylation Sites: 1 | Go to GlyGen: Q86WC4-1 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| H(+)/Cl(-) exchange transporter 7 | 825 | Homo sapiens | Mutation(s): 0 Gene Names: CLCN7 Membrane Entity: Yes | 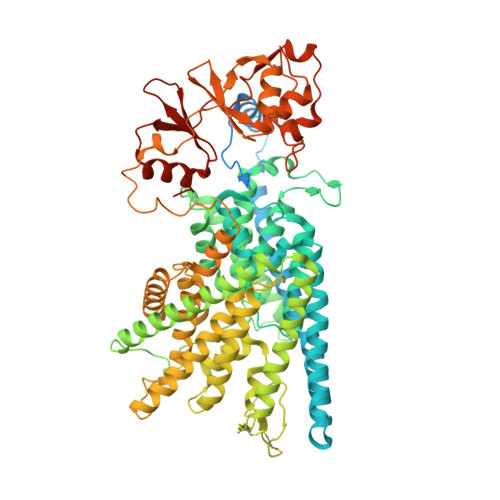 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P51798 GTEx: ENSG00000103249 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P51798 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| ATP Download:Ideal Coordinates CCD File | G [auth C], J [auth D] | ADENOSINE-5'-TRIPHOSPHATE C10 H16 N5 O13 P3 ZKHQWZAMYRWXGA-KQYNXXCUSA-N |  | ||
| CL Download:Ideal Coordinates CCD File | H [auth C], I [auth C], K [auth D], L [auth D] | CHLORIDE ION Cl VEXZGXHMUGYJMC-UHFFFAOYSA-M |  | ||
| Task | Software Package | Version |
|---|---|---|
| MODEL REFINEMENT | PHENIX | |