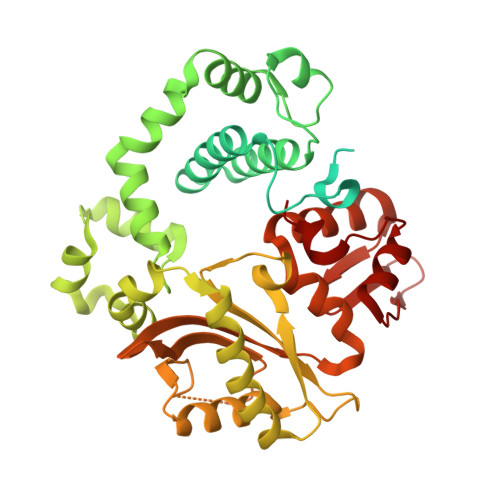
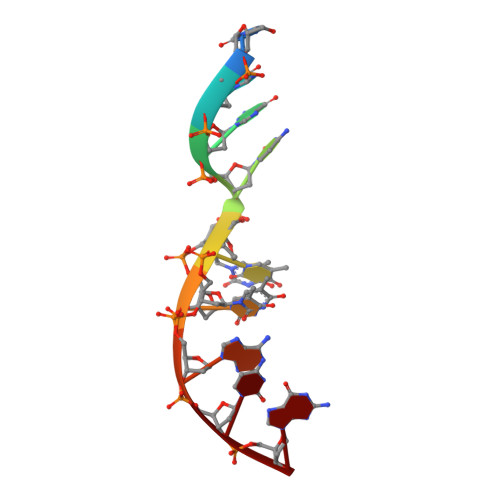
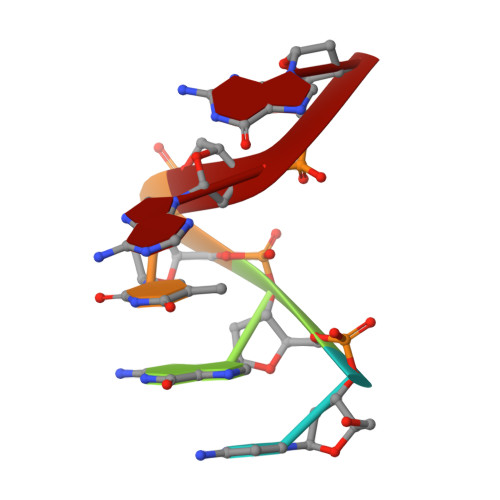
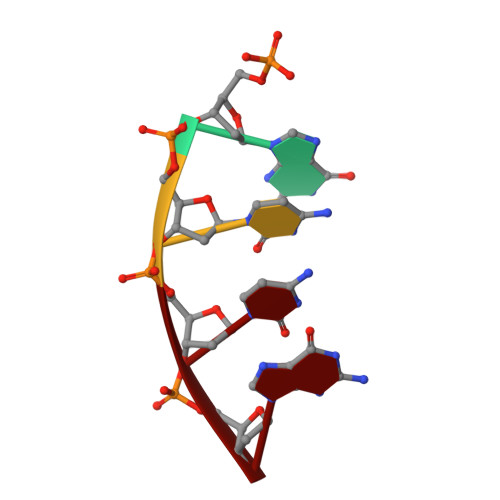
Mechanism of genome instability mediated by human DNA polymerase mu misincorporation.
Guo, M., Wang, Y., Tang, Y., Chen, Z., Hou, J., Dai, J., Wang, Y., Wang, L., Xu, H., Tian, B., Hua, Y., Zhao, Y.(2021) Nat Commun 12: 3759-3759
- PubMed: 34145298 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-24096-7
- Primary Citation Related Structures:
7CO6, 7CO8, 7CO9, 7COA, 7COB, 7COC, 7COD - PubMed Abstract:
Pol μ is capable of performing gap-filling repair synthesis in the nonhomologous end joining (NHEJ) pathway. Together with DNA ligase, misincorporation of dGTP opposite the templating T by Pol μ results in a promutagenic T:G mispair, leading to genomic instability. Here, crystal structures and kinetics of Pol μ substituting dGTP for dATP on gapped DNA substrates containing templating T were determined and compared. Pol μ is highly mutagenic on a 2-nt gapped DNA substrate, with T:dGTP base pairing at the 3' end of the gap. Two residues (Lys438 and Gln441) interact with T:dGTP and fine tune the active site microenvironments. The in-crystal misincorporation reaction of Pol μ revealed an unexpected second dGTP in the active site, suggesting its potential mutagenic role among human X family polymerases in NHEJ.
- Institute of Biophysics, College of Life Sciences, Zhejiang University, Hangzhou, Zhejiang, China.
Organizational Affiliation: