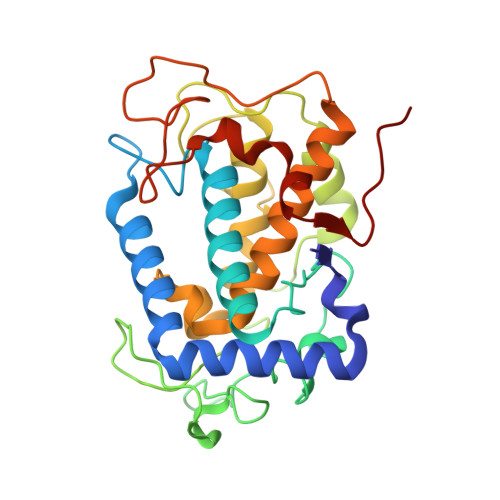
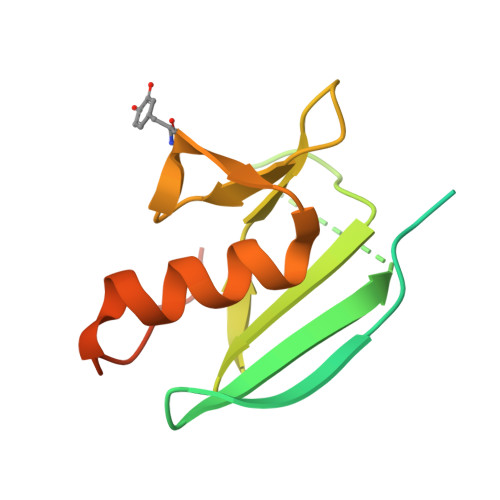
The basicity of an active-site water molecule discriminates between tyrosinase and catechol oxidase activity.
Matoba, Y., Oda, K., Muraki, Y., Masuda, T.(2021) Int J Biol Macromol 183: 1861-1870
- PubMed: 34089758 Search on PubMed
- DOI: https://doi.org/10.1016/j.ijbiomac.2021.05.206
- Primary Citation Related Structures:
7CIT, 7CIY - PubMed Abstract:
Tyrosinase (Ty) and catechol oxidase (CO) are members of type-3 copper enzymes. While Ty catalyzes both phenolase and catecholase reactions, CO catalyzes only the latter reaction. In the present study, Ty was found to catalyze the catecholase reaction, but hardly the phenolase reaction in the presence of the metallochaperon called "caddie protein (Cad)". The ability of the substrates to dissociate the motif shielding the active-site pocket seems to contribute critically to the substrate specificity of Ty. In addition, a mutation at the N191 residue, which forms a hydrogen bond with a water molecule near the active center, decreased the inherent ratio of phenolase versus catecholase activity. Unlike the wild-type complex, reaction intermediates were not observed when the catalytic reaction toward the Y98 residue of Cad was progressed in the crystalline state. The increased basicity of the water molecule may be necessary to inhibit the proton transfer from the conjugate acid to a hydroxide ion bridging the two copper ions. The deprotonation of the substrate hydroxyl by the bridging hydroxide seems to be significant for the efficient catalytic cycle of the phenolase reaction.
- Faculty of Pharmacy, Yasuda Women's University, Yasuhigashi 6-13-1, Asaminami-ku, Hiroshima, 731-0153, Japan. Electronic address: matoba@yasuda-u.ac.jp.
Organizational Affiliation: