Structural Study on Anti-CRISPR Protein AcrVA2
Chen, P., Sun, W., Cheng, Z., Yang, J., Wang, M., Wang, J., Chen, H., Liu, L., Wang, Y.(2021) PROG BIOCHEM BIOPHYS
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| AcrVA2 | 323 | Moraxella bovoculi | Mutation(s): 0 | 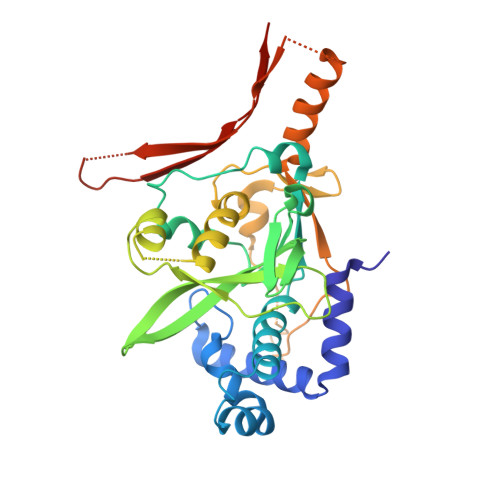 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | A0A7M4DUH0 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| SPD Download:Ideal Coordinates CCD File | C [auth A], K [auth A], L [auth B] | SPERMIDINE C7 H19 N3 ATHGHQPFGPMSJY-UHFFFAOYSA-N |  | ||
| EDO Download:Ideal Coordinates CCD File | D [auth A] E [auth A] F [auth A] G [auth A] H [auth A] | 1,2-ETHANEDIOL C2 H6 O2 LYCAIKOWRPUZTN-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 84.287 | α = 90 |
| b = 84.287 | β = 90 |
| c = 264.171 | γ = 120 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| HKL-2000 | data reduction |
| HKL-2000 | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Natural Science Foundation of China (NSFC) | China | 91440201 |
| National Natural Science Foundation of China (NSFC) | China | 31700662 |