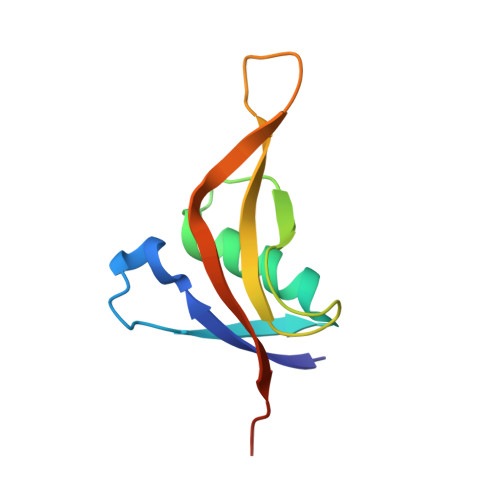
A 1.3 angstrom high-resolution crystal structure of an anti-CRISPR protein, AcrI E2.
Lee, S.Y., Kim, G.E., Kim, Y.G., Park, H.H.(2020) Biochem Biophys Res Commun 533: 751-757
- PubMed: 32988588 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2020.09.067
- Primary Citation Related Structures:
7CHQ - PubMed Abstract:
As a result of bacterial infection with viruses, bacteria have developed CRISPR-Cas as an adaptive immune system, which allows them to destroy the viral genetic material introduced via infection. However, viruses have also evolved to develop multiple anti-CRISPR proteins, which are capable of inactivating the CRISPR-Cas adaptive immune system to combat bacteria. In this study, we aimed to elucidate the molecular mechanisms associated with anti-CRISPR proteins by determining a high-resolution crystal structure (1.3 Å) of Type I-E anti-CRISPR protein called AcrIE2. Our structural analysis revealed that AcrIE2 was composed of unique folds comprising five antiparallel β-sheets (β1∼β5) surrounding one α-helix (α1) in the order, β 2 β 1 α 1 β 5 β 4 β 3 . Structural comparison of AcrIE2 with a structural homolog called AcrIF9 showed that AcrIE2 contained a long and flexible β4-β5 connecting loop and a distinct surface feature. These results indicated that the inhibitory mechanism of AcrIE2 might be different from that of AcrIF9. This unique structure of AcrIE2 indicates its special mode of CRISPR-Cas inhibitory activity. Therefore, this study helps us understand the diversity in the inhibitory mechanisms of Acr family.
- Department of Global Innovative Drugs, Graduate School of Chung-Ang University, Seoul, 06974, Republic of Korea; College of Pharmacy, Chung-Ang University, Seoul, 06974, Republic of Korea.
Organizational Affiliation: