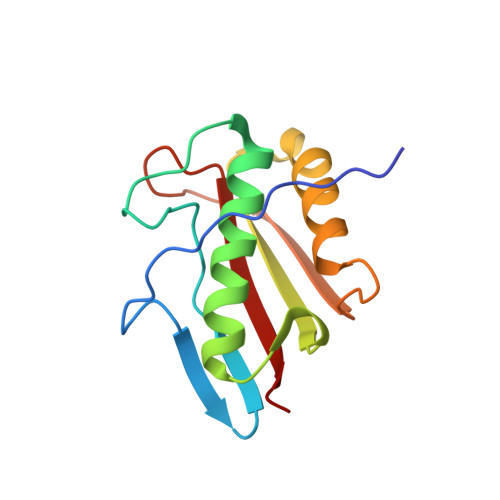
Tetramer formation of Bacillus subtilis YabJ protein that belongs to YjgF/YER057c/UK114 family.
Fujimoto, Z., Hong, L.T.T., Kishine, N., Suzuki, N., Kimura, K.(2021) Biosci Biotechnol Biochem 85: 297-306
- PubMed: 33590041 Search on PubMed
- DOI: https://doi.org/10.1093/bbb/zbaa037
- Primary Citation Related Structures:
5Y6U, 7CD2, 7CD3, 7CD4 - PubMed Abstract:
Bacillus subtilis YabJ protein belongs to the highly conserved YjgF/YER057c/UK114 family, which has a homotrimeric quaternary structure. The dominant allele of yabJ gene that is caused by a single amino acid mutation of Ser103Phe enables poly-γ-glutamic acid (γPGA) production of B. subtilis under conditions where the cell-density signal transduction was disturbed by the loss of DegQ function. X-ray crystallography of recombinant proteins revealed that unlike the homotrimeric wild-type YabJ, the mutant YabJ(Ser103Phe) had a homotetrameric quaternary structure, and the structural change appeared to be triggered by an inversion of the fifth β-strand. The YabJ homotetramer has a hole that is highly accessible, penetrating through the tetramer, and 2 surface concaves as potential ligand-binding sites. Western blot analyses revealed that the conformational change was also induced in vivo by the Ser103Phe mutation.
- Advanced Analysis Center, National Agriculture and Food Research Organization (NAAC/NARO), Tsukuba, Ibaraki, Japan.
Organizational Affiliation: