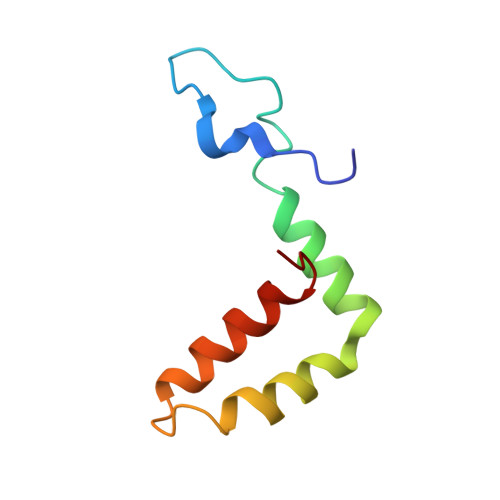
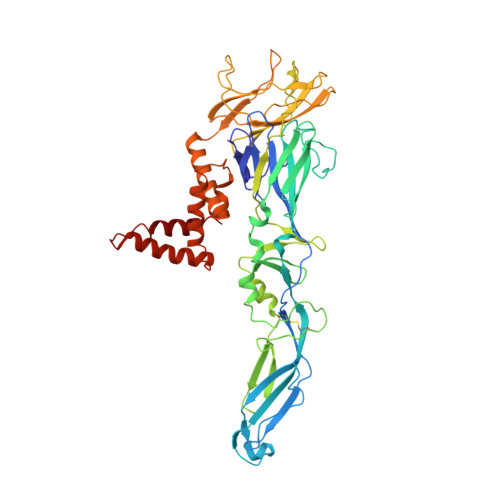
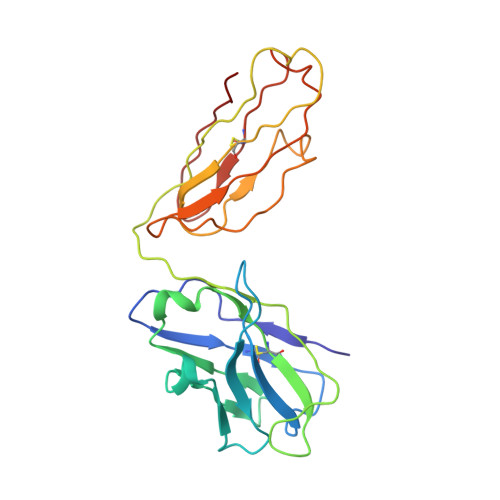
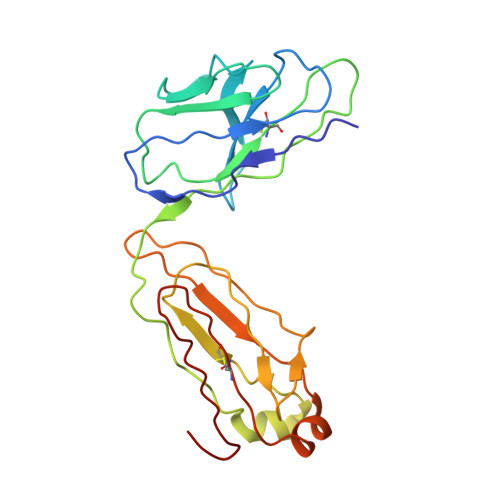
A complex between the Zika virion and the Fab of a broadly cross-reactive neutralizing monoclonal antibody revealed by cryo-EM and single particle analysis at 4.1 angstrom resolution.
Tyagi, A., Ahmed, T., Shi, J., Bhushan, S.(2020) J Struct Biol X 4: 100028-100028
- PubMed: 32647830 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.yjsbx.2020.100028
- Primary Citation Related Structures:
7CBP - PubMed Abstract:
Zika virus (ZIKV) recently emerged as a major public health concern because it can cause fetal microcephaly and neurological disease such as the Guillain-Barré syndrome. A particularly potent class of broadly neutralizing antibodies (nAbs) targets a quaternary epitope located at the interface of two envelope proteins monomers, exposed at the surface of the mature virion. This "E-dimer-dependent epitope" (EDE), comprises the fusion loop of one monomer at the tip of domain II of E and a portion of the domains I and III of the adjacent monomer. Since this epitope largely overlaps with the binding site of the precursor membrane protein (prM) during Zika virion maturation, its molecular surface is evolutionary conserved in flaviviruses such as Dengue and Zika viruses, and can elicit antibodies that broadly neutralize various ZIKV strains. Here, we present a cryo-EM reconstruction at 4.1 Å resolution of the virion bound to the antigen binding fragment (Fab) of an antibody that targets this mutationally-constrained quaternary epitope. The Fab incompletely covers the surface of the virion as it does not bind next to its 5-fold icosahedral axes. The structure reveals details of the binding mode of this potent neutralizing class of antibodies and can inform the design of immunogens and vaccines targeting this conserved epitope.
- School of Biological Sciences, Nanyang Technological University, Singapore.
Organizational Affiliation: