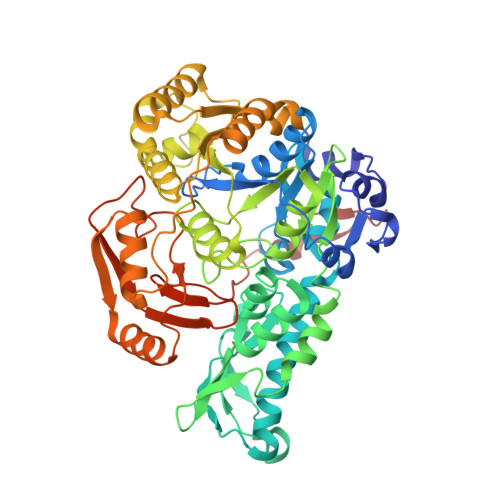
Biochemical properties and crystal structure of formate-tetrahydrofolate ligase from Methylobacterium extorquens CM4.
Kim, S., Lee, S.H., Seo, H., Kim, K.J.(2020) Biochem Biophys Res Commun 528: 426-431
- PubMed: 32505353 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2020.05.198
- Primary Citation Related Structures:
7C11 - PubMed Abstract:
Methylobacterium extorquens is a methylotroph model organism that has the ability to assimilate formate using the tetrahydrofolate (THF) pathway. The formate-tetrahydrofolate ligase from M. extorquens (MeFtfL) is an enzyme involved in the THF pathway that catalyzes the conversion of formate, THF, and ATP into formyltetrahydrofolate and ADP. To investigate the biochemical properties of MeFtfL, we evaluated the metal usage and enzyme kinetics of the enzyme. MeFtfL uses the Mg ion for catalytic activity, but also has activity for Mn and Ca ions. The enzyme kinetics analysis revealed that K m value of farmate was much higher than THF and ATP, which shows that the ligation activity of MeFtfL is highly dependent on formation concentration. We also determined the crystal structure of MeFtfL at 2.8 Å resolution. MeFtfL functions as a tetramer, and each monomer consists of three domains. The structural superposition of MeFtfL with FtfL from Moorella thermoacetica allowed us to predict the substrate binding site of the enzyme.
- School of Life Sciences, KNU Creative BioResearch Group, Kyungpook National University, Daegu, 41566, Republic of Korea; KNU Institute for Microorganisms, Kyungpook National University, Daegu, 41566, Republic of Korea.
Organizational Affiliation: