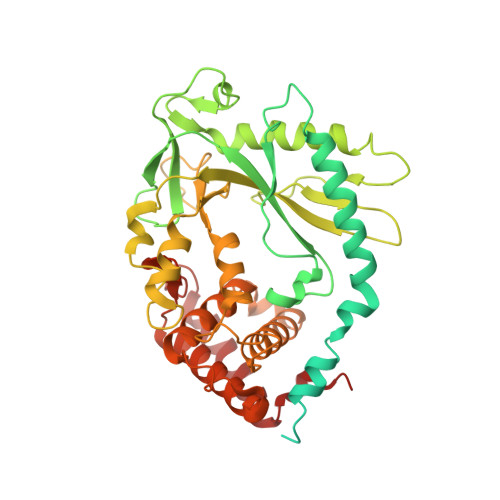
Mn2+Directly Activates cGAS and Structural Analysis Suggests Mn2+Induces a Noncanonical Catalytic Synthesis of 2'3'-cGAMP.
Zhao, Z., Ma, Z., Wang, B., Guan, Y., Su, X.D., Jiang, Z.(2020) Cell Rep 32: 108053-108053
- PubMed: 32814054 Search on PubMed
- DOI: https://doi.org/10.1016/j.celrep.2020.108053
- Primary Citation Related Structures:
7BUJ, 7BUM, 7BUQ - PubMed Abstract:
DNA binding allosterically activates the cytosolic DNA sensor cGAS (cyclic GMP-AMP [cGAMP] synthase) to synthesize 2'3'-cGAMP, using Mg 2+ as the metal cofactor that catalyzes two nucleotidyl-transferring reactions. We previously found that Mn 2+ potentiates cGAS activation, but the underlying mechanism remains unclear. Here, we report that Mn 2+ directly activates cGAS. Structural analysis reveals that Mn 2+ -activated cGAS undergoes globally similar conformational changes to DNA-activated cGAS but forms a unique η1 helix to widen the catalytic pocket, allowing substrate entry and cGAMP synthesis. Strikingly, in Mn 2+ -activated cGAS, the linear intermediates pppGpG and pGpA take an inverted orientation in the active pocket, suggesting a noncanonical but accelerated cGAMP cyclization without substrate flip-over. Moreover, unlike the octahedral coordination around Mg 2+ , the two catalytic Mn 2+ are coordinated by triphosphate moiety of the inverted substrate, independent of the catalytic triad residues. Our findings thus uncover Mn 2+ as a cGAS activator that initiates noncanonical 2'3'-cGAMP synthesis.
- Key Laboratory of Cell Proliferation and Differentiation of the Ministry of Education, School of Life Sciences, Peking University, Beijing 100871, China; Peking-Tsinghua Center for Life Sciences, Peking University, Beijing 100871, China.
Organizational Affiliation: