SLX4 control MUS81 endonuclease function in telomere through an intermolecular SAP domain
Wan, B., Wu, J., Lei, M.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| MUS81 endonuclease homolog (Yeast), isoform CRA_b | 98 | Homo sapiens | Mutation(s): 0 Gene Names: MUS81, hCG_23303 EC: 3.1.22 | 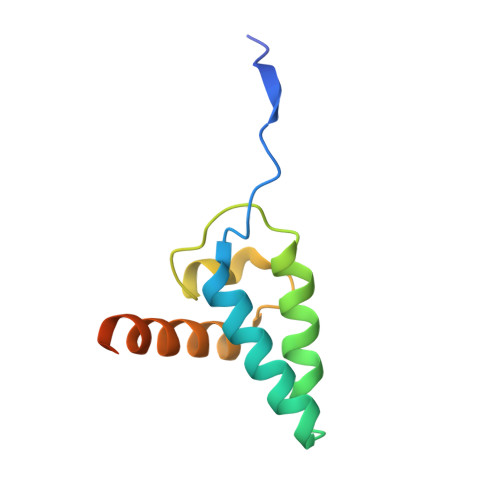 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q96NY9 GTEx: ENSG00000172732 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q96NY9 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Structure-specific endonuclease subunit SLX4 | 61 | Homo sapiens | Mutation(s): 0 Gene Names: SLX4, BTBD12, KIAA1784, KIAA1987 | 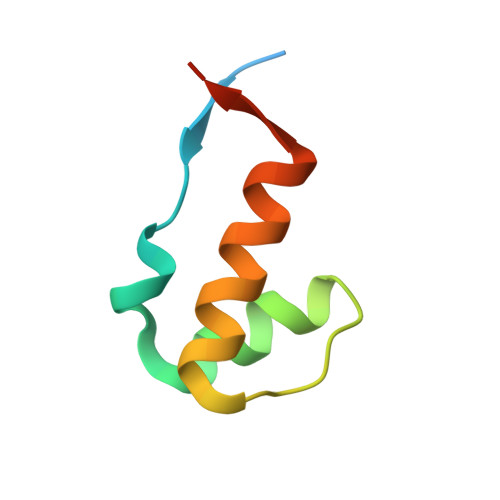 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q8IY92 GTEx: ENSG00000188827 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q8IY92 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| GLY Download:Ideal Coordinates CCD File | C [auth A], D [auth A] | GLYCINE C2 H5 N O2 DHMQDGOQFOQNFH-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 59.118 | α = 90 |
| b = 59.118 | β = 90 |
| c = 86.425 | γ = 120 |
| Software Name | Purpose |
|---|---|
| SCALEPACK | data scaling |
| REFMAC | refinement |
| PDB_EXTRACT | data extraction |
| Coot | model building |
| PHENIX | phasing |