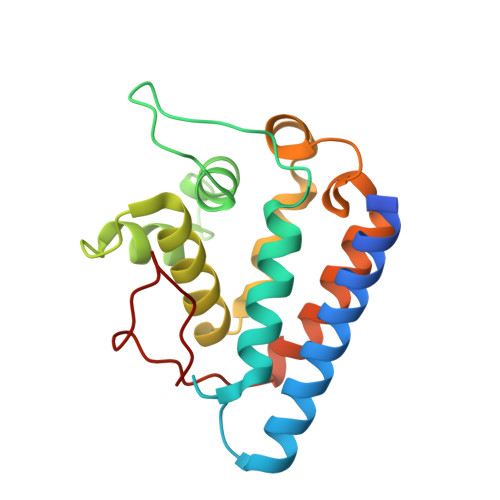
Crystal structure of the FYCO1 RUN domain suggests possible interfaces with small GTPases.
Sakurai, S., Shimizu, T., Ohto, U.(2020) Acta Crystallogr F Struct Biol Commun 76: 326-333
- PubMed: 32744243 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X20009012
- Primary Citation Related Structures:
7BQI - PubMed Abstract:
FYCO1 is a multidomain adaptor protein that plays an important role in autophagy by mediating the kinesin-dependent microtubule plus-end-directed transport of autophagosomes. FYCO1 contains a RUN domain, which is hypothesized to function as a specific effector for members of the Ras superfamily of small GTPases, but its role has not been well characterized and its interaction partner(s) have not been identified. Here, the crystal structure of the FYCO1 RUN domain was determined at 1.3 Å resolution. The overall structure of the FYCO1 RUN domain was similar to those of previously reported RUN domains. Detailed structural comparisons with other RUN domains and docking studies suggested a possible interaction interface of the FYCO1 RUN domain with small GTPases of the Ras superfamily.
- Graduate School of Pharmaceutical Sciences, The University of Tokyo, Hongo, Bunkyo-ku, Tokyo 113-0033, Japan.
Organizational Affiliation: