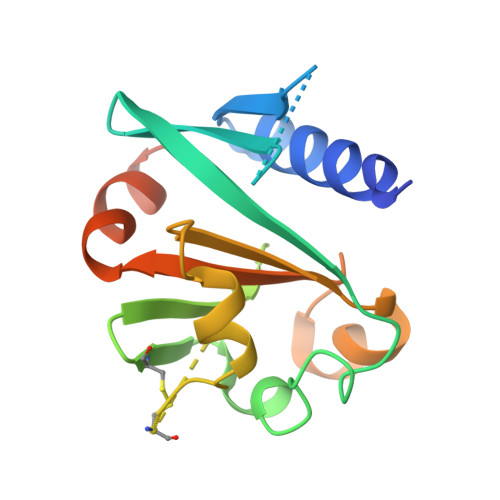
Crystal structure and initial characterization of a novel archaeal-like Holliday junction-resolving enzyme from Thermus thermophilus phage Tth15-6.
Ahlqvist, J., Linares-Pasten, J.A., Hakansson, M., Jasilionis, A., Kwiatkowska-Semrau, K., Friðjonsson, O.H., Kaczorowska, A.K., Dabrowski, S., Aevarsson, A., Hreggviðsson, G.O., Al-Karadaghi, S., Kaczorowski, T., Nordberg Karlsson, E.(2022) Acta Crystallogr D Struct Biol 78: 212-227
- PubMed: 35102887 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2059798321012298
- Primary Citation Related Structures:
7BGS, 7BNX - PubMed Abstract:
This study describes the production, characterization and structure determination of a novel Holliday junction-resolving enzyme. The enzyme, termed Hjc_15-6, is encoded in the genome of phage Tth15-6, which infects Thermus thermophilus. Hjc_15-6 was heterologously produced in Escherichia coli and high yields of soluble and biologically active recombinant enzyme were obtained in both complex and defined media. Amino-acid sequence and structure comparison suggested that the enzyme belongs to a group of enzymes classified as archaeal Holliday junction-resolving enzymes, which are typically divalent metal ion-binding dimers that are able to cleave X-shaped dsDNA-Holliday junctions (Hjs). The crystal structure of Hjc_15-6 was determined to 2.5 Å resolution using the selenomethionine single-wavelength anomalous dispersion method. To our knowledge, this is the first crystal structure of an Hj-resolving enzyme originating from a bacteriophage that can be classified as an archaeal type of Hj-resolving enzyme. As such, it represents a new fold for Hj-resolving enzymes from phages. Characterization of the structure of Hjc_15-6 suggests that it may form a dimer, or even a homodimer of dimers, and activity studies show endonuclease activity towards Hjs. Furthermore, based on sequence analysis it is proposed that Hjc_15-6 has a three-part catalytic motif corresponding to E-SD-EVK, and this motif may be common among other Hj-resolving enzymes originating from thermophilic bacteriophages.
- Biotechnology, Department of Chemistry, Lund University, PO Box 124, 221 00 Lund, Sweden.
Organizational Affiliation: