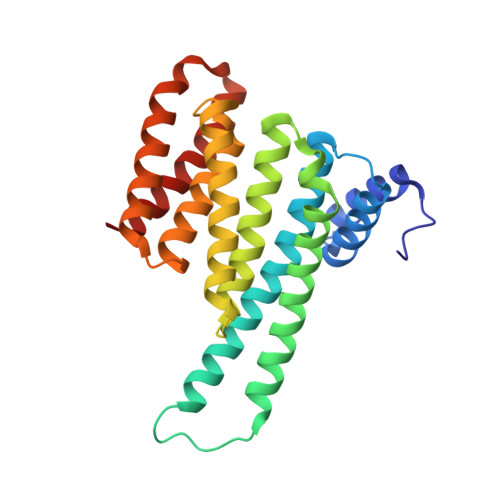
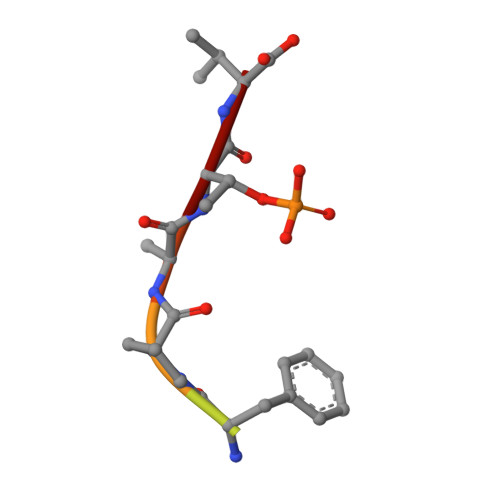
Exploration of a 14-3-3 PPI Pocket by Covalent Fragments as Stabilizers.
Sijbesma, E., Hallenbeck, K.K., Andrei, S.A., Rust, R.R., Adriaans, J.M.C., Brunsveld, L., Arkin, M.R., Ottmann, C.(2021) ACS Med Chem Lett 12: 976-982
- PubMed: 34136078 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsmedchemlett.1c00088
- Primary Citation Related Structures:
7B9M, 7B9R, 7B9T, 7BA3, 7BA5, 7BA6, 7BA7, 7BA8, 7BA9, 7BAA, 7BAB - PubMed Abstract:
The systematic discovery of functional fragments binding to the composite interface of protein complexes is a first critical step for the development of orthosteric stabilizers of protein-protein interactions (PPIs). We have previously shown that disulfide trapping successfully yielded covalent stabilizers for the PPI of 14-3-3 with the estrogen receptor ERα. Here we provide an assessment of the composite PPI target pocket and the molecular characteristics of various fragments binding to a specific subpocket. Evaluating structure-activity relationships highlights the basic principles for PPI stabilization by these covalent fragments that engage a relatively large and exposed binding pocket at the protein/peptide interface with a "molecular glue" mode of action.
- Laboratory of Chemical Biology, Department of Biomedical Engineering and Institute for Complex Molecular Systems (ICMS), Eindhoven University of Technology, 5600 MB Eindhoven, The Netherlands.
Organizational Affiliation: