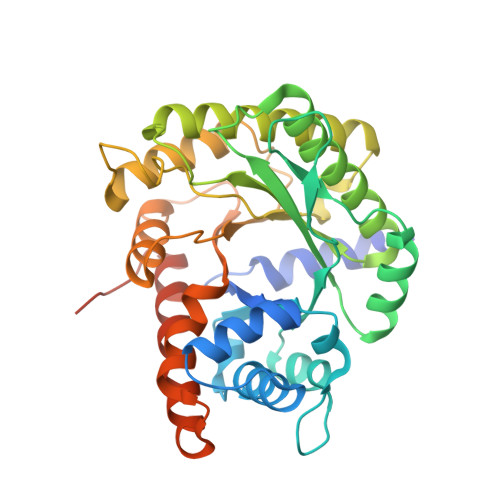
Crystal structure of chloroplast fructose-1,6-bisphosphate aldolase from the green alga Chlamydomonas reinhardtii.
Le Moigne, T., Sarti, E., Nourisson, A., Zaffagnini, M., Carbone, A., Lemaire, S.D., Henri, J.(2022) J Struct Biol 214: 107873-107873
- PubMed: 35680033 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2022.107873
- Primary Citation Related Structures:
7B2N - PubMed Abstract:
The Calvin-Benson cycle fixes carbon dioxide into organic triosephosphates through the collective action of eleven conserved enzymes. Regeneration of ribulose-1,5-bisphosphate, the substrate of Rubisco-mediated carboxylation, requires two lyase reactions catalyzed by fructose-1,6-bisphosphate aldolase (FBA). While cytoplasmic FBA has been extensively studied in non-photosynthetic organisms, functional and structural details are limited for chloroplast FBA encoded by oxygenic phototrophs. Here we determined the crystal structure of plastidial FBA from the unicellular green alga Chlamydomonas reinhardtii (Cr). We confirm that CrFBA folds as a TIM barrel, describe its catalytic pocket and homo-tetrameric state. Multiple sequence profiling classified the photosynthetic paralogs of FBA in a distinct group from non-photosynthetic paralogs. We mapped the sites of thiol- and phospho-based post-translational modifications known from photosynthetic organisms and predict their effects on enzyme catalysis.
- Sorbonne Université, CNRS, UMR 7238, Institut de Biologie Paris-Seine, Laboratoire de Biologie Computationnelle et Quantitative, 4 place Jussieu, F-75005 Paris, France; Sorbonne Université, CNRS, UMR 8226, Institut de Biologie Physico-Chimique, Laboratoire de Biologie Moléculaire et Cellulaire des Eucaryotes, 13 rue Pierre et Marie Curie, 75005 Paris, France; Faculty of Sciences, Doctoral School of Plant Sciences, Université Paris-Saclay, 91190 Saint-Aubin, France.
Organizational Affiliation: