Carrier Protein-Free Enzymatic Biaryl Coupling in Arylomycin A2 Assembly and Structure of the Cytochrome P450 AryC.
Aldemir, H., Shu, S., Schaefers, F., Hong, H., Richarz, R., Harteis, S., Einsiedler, M., Milzarek, T.M., Schneider, S., Gulder, T.A.M.(2022) Chemistry 28: e202103389-e202103389
- PubMed: 34725865 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/chem.202103389
- Primary Citation Related Structures:
7AYX - PubMed Abstract:
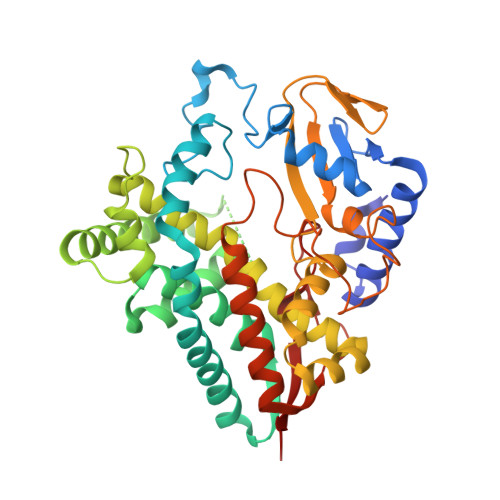
The arylomycin antibiotics are potent inhibitors of bacterial type I signal peptidase. These lipohexapeptides contain a biaryl structural motif reminiscent of glycopeptide antibiotics. We herein describe the functional and structural evaluation of AryC, the cytochrome P450 performing biaryl coupling in biosynthetic arylomycin assembly. Unlike its enzymatic counterparts in glycopeptide biosynthesis, AryC converts free substrates without the requirement of any protein interaction partner, likely enabled by a strongly hydrophobic cavity at the surface of AryC pointing to the substrate tunnel. This activity enables chemo-enzymatic assembly of arylomycin A2 that combines the advantages of liquid- and solid-phase peptide synthesis with late-stage enzymatic cross-coupling. The reactivity of AryC is unprecedented in cytochrome P450-mediated biaryl construction in non-ribosomal peptides, in which peptidyl carrier protein (PCP)-tethering so far was shown crucial both in vivo and in vitro.
- Chair of Technical Biochemistry, Technical University of Dresden, Bergstraße 66, 01069, Dresden, Germany.
Organizational Affiliation: