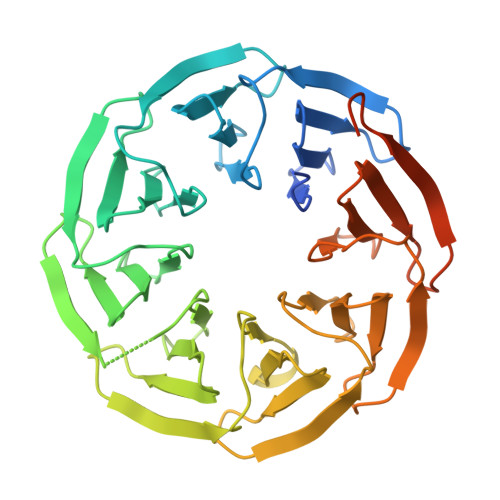
Crystal structures of Scone: pseudosymmetric folding of a symmetric designer protein.
Mylemans, B., Killian, T., Vandebroek, L., Van Meervelt, L., Tame, J.R.H., Parac-Vogt, T.N., Voet, A.R.D.(2021) Acta Crystallogr D Struct Biol 77: 933-942
- PubMed: 34196619 Search on PubMed
- DOI: https://doi.org/10.1107/S2059798321005787
- Primary Citation Related Structures:
7AWY, 7AWZ, 7AX0, 7AX2 - PubMed Abstract:
Recent years have seen an increase in the development of computational proteins, including symmetric ones. A ninefold-symmetric β-propeller protein named Cake has recently been developed. Here, attempts were made to further engineer this protein into a threefold-symmetric nine-bladed propeller using computational design. Two nine-bladed propeller proteins were designed, named Scone-E and Scone-R. Crystallography, however, revealed the structure of both designs to adopt an eightfold conformation with distorted termini, leading to a pseudo-symmetric protein. One of the proteins could only be crystallized upon the addition of a polyoxometalate, highlighting the usefulness of these molecules as crystallization additives.
- Laboratory of Biomolecular Modelling and Design, Department of Chemistry, KU Leuven, Celestijnenlaan 200G, 3001 Leuven, Vlaams Brabant, Belgium.
Organizational Affiliation: