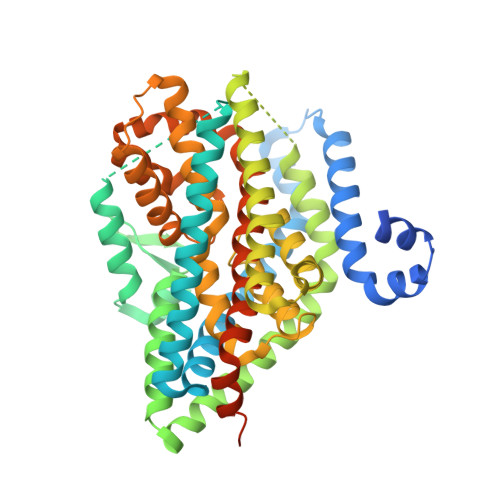
The ion-coupling mechanism of human excitatory amino acid transporters.
Canul-Tec, J.C., Kumar, A., Dhenin, J., Assal, R., Legrand, P., Rey, M., Chamot-Rooke, J., Reyes, N.(2022) EMBO J 41: e108341-e108341
- PubMed: 34747040 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.15252/embj.2021108341
- Primary Citation Related Structures:
7AWL, 7AWM, 7AWN, 7AWP, 7AWQ, 7NPW - PubMed Abstract:
Excitatory amino acid transporters (EAATs) maintain glutamate gradients in the brain essential for neurotransmission and to prevent neuronal death. They use ionic gradients as energy source and co-transport transmitter into the cytoplasm with Na + and H + , while counter-transporting K + to re-initiate the transport cycle. However, the molecular mechanisms underlying ion-coupled transport remain incompletely understood. Here, we present 3D X-ray crystallographic and cryo-EM structures, as well as thermodynamic analysis of human EAAT1 in different ion bound conformations, including elusive counter-transport ion bound states. Binding energies of Na + and H + , and unexpectedly Ca 2+ , are coupled to neurotransmitter binding. Ca 2+ competes for a conserved Na + site, suggesting a regulatory role for Ca 2+ in glutamate transport at the synapse, while H + binds to a conserved glutamate residue stabilizing substrate occlusion. The counter-transported ion binding site overlaps with that of glutamate, revealing the K + -based mechanism to exclude the transmitter during the transport cycle and to prevent its neurotoxic release on the extracellular side.
- Membrane Protein Mechanisms Unit, Institut Pasteur, Paris, France.
Organizational Affiliation: