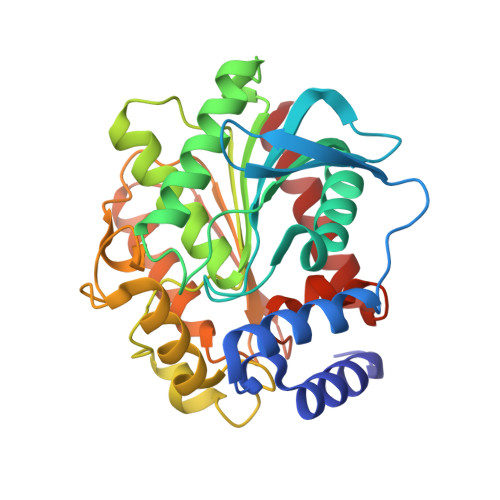
Biochemical and Structural Characterization of a novel thermophilic esterase EstD11 provide catalytic insights for the HSL family.
Miguel-Ruano, V., Rivera, I., Rajkovic, J., Knapik, K., Torrado, A., Otero, J.M., Beneventi, E., Becerra, M., Sanchez-Costa, M., Hidalgo, A., Berenguer, J., Gonzalez-Siso, M.I., Cruces, J., Rua, M.L., Hermoso, J.A.(2021) Comput Struct Biotechnol J 19: 1214-1232
- PubMed: 33680362 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.csbj.2021.01.047
- Primary Citation Related Structures:
7AT0, 7AT2, 7AT3, 7AT4, 7ATD, 7ATF, 7ATQ, 7AUY, 7AV5, 7NB5 - PubMed Abstract:
A novel esterase, EstD11, has been discovered in a hot spring metagenomic library. It is a thermophilic and thermostable esterase with an optimum temperature of 60°C. A detailed substrate preference analysis of EstD11 was done using a library of chromogenic ester substrate that revealed the broad substrate specificity of EstD11 with significant measurable activity against 16 substrates with varied chain length, steric hindrance, aromaticity and flexibility of the linker between the carboxyl and the alcohol moiety of the ester. The tridimensional structures of EstD11 and the inactive mutant have been determined at atomic resolutions. Structural and bioinformatic analysis, confirm that EstD11 belongs to the family IV, the hormone-sensitive lipase (HSL) family, from the α/β-hydrolase superfamily. The canonical α/β-hydrolase domain is completed by a cap domain, composed by two subdomains that can unmask of the active site to allow the substrate to enter. Eight crystallographic complexes were solved with different substrates and reaction products that allowed identification of the hot-spots in the active site underlying the specificity of the protein. Crystallization and/or incubation of EstD11 at high temperature provided unique information on cap dynamics and a first glimpse of enzymatic activity in vivo . Very interestingly, we have discovered a unique Met zipper lining the active site and the cap domains that could be essential in pivotal aspects as thermo-stability and substrate promiscuity in EstD11.
- Department of Crystallography and Structural Biology, Institute of Physical-Chemistry "Rocasolano", Spanish National Research Council (CSIC), Madrid, Spain.
Organizational Affiliation: