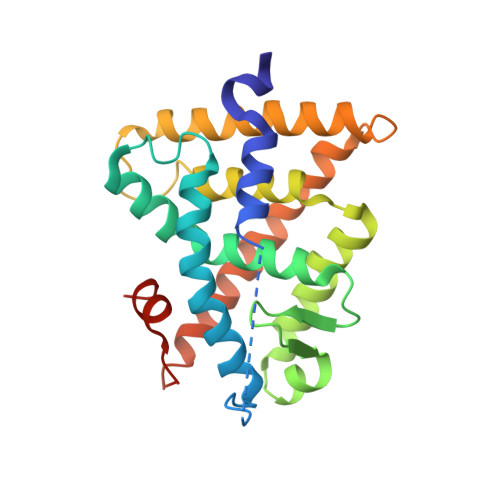
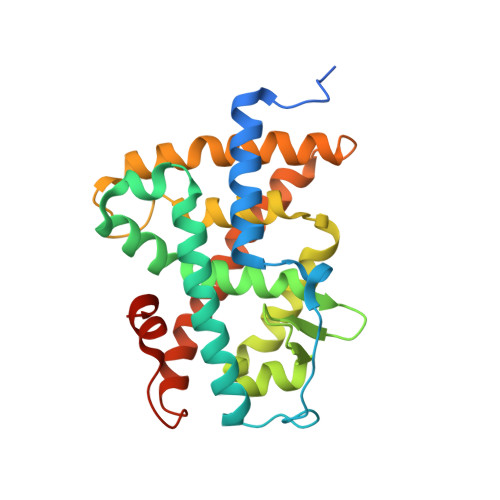
Structural Insights into the Interaction of the Intrinsically Disordered Co-activator TIF2 with Retinoic Acid Receptor Heterodimer (RXR/RAR).
Senicourt, L., le Maire, A., Allemand, F., Carvalho, J.E., Guee, L., Germain, P., Schubert, M., Bernado, P., Bourguet, W., Sibille, N.(2021) J Mol Biology 433: 166899-166899
- PubMed: 33647291 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2021.166899
- Primary Citation Related Structures:
7AOS, 7APO, 7BK4 - PubMed Abstract:
Retinoic acid receptors (RARs) and retinoid X receptors (RXRs) form heterodimers that activate target gene transcription by recruiting co-activator complexes in response to ligand binding. The nuclear receptor (NR) co-activator TIF2 mediates this recruitment by interacting with the ligand-binding domain (LBD) of NRs trough the nuclear receptor interaction domain (TIF2 NRID ) containing three highly conserved α-helical LxxLL motifs (NR-boxes). The precise binding mode of this domain to RXR/RAR is not clear due to the disordered nature of TIF2. Here we present the structural characterization of TIF2 NRID by integrating several experimental (NMR, SAXS, Far-UV CD, SEC-MALS) and computational data. Collectively, the data are in agreement with a largely disordered protein with partially structured regions, including the NR-boxes and their flanking regions, which are evolutionary conserved. NMR and X-ray crystallographic data on TIF2 NRID in complex with RXR/RAR reveal a multisite binding of the three NR-boxes as well as an active role of their flanking regions in the interaction.
- Centre de Biochimie Structurale (CBS), CNRS, INSERM, Univ Montpellier, Montpellier, France.
Organizational Affiliation: