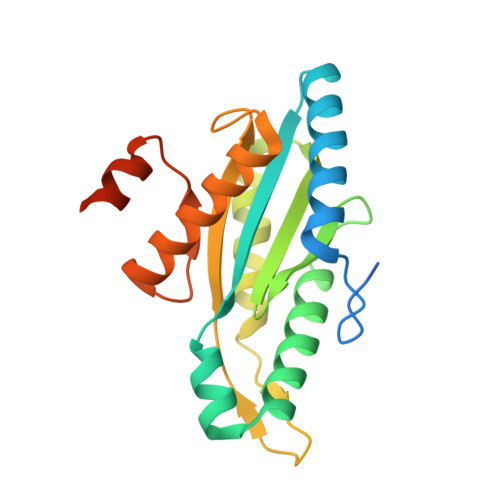
Studying GGDEF Domain in the Act: Minimize Conformational Frustration to Prevent Artefacts.
Mantoni, F., Scribani Rossi, C., Paiardini, A., Di Matteo, A., Cappellacci, L., Petrelli, R., Ricciutelli, M., Paone, A., Cutruzzola, F., Giardina, G., Rinaldo, S.(2021) Life (Basel) 11
- PubMed: 33418960 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/life11010031
- Primary Citation Related Structures:
7A7E - PubMed Abstract:
GGDEF-containing proteins respond to different environmental cues to finely modulate cyclic diguanylate (c-di-GMP) levels in time and space, making the allosteric control a distinctive trait of the corresponding proteins. The diguanylate cyclase mechanism is emblematic of this control: two GGDEF domains, each binding one GTP molecule, must dimerize to enter catalysis and yield c-di-GMP. The need for dimerization makes the GGDEF domain an ideal conformational switch in multidomain proteins. A re-evaluation of the kinetic profile of previously characterized GGDEF domains indicated that they are also able to convert GTP to GMP: this unexpected reactivity occurs when conformational issues hamper the cyclase activity. These results create new questions regarding the characterization and engineering of these proteins for in solution or structural studies.
- Department of Biochemical Sciences "A. Rossi Fanelli", Sapienza University of Rome, Piazzale Aldo Moro, 5, 00185 Rome, Italy.
Organizational Affiliation: